Model Service
Model Service Overview
Model service is key to achieving a full lifecycle closed-loop in MLOps. It supports real-time prediction and online inference by one-click release of models from model management as API services, and provides complete service monitoring and operations capacity. In addition, model service is deeply integrated with other core modules of data science, supporting one-click release of models as standard REST APIs and offering multidimensional traffic, resource monitoring, and running log backtracking capacity.
Function Operation Guide
WeData's model service provides features such as create service, service monitoring, and debug to manage services more efficiently and ensure their validity in a production environment.
Creating a Model Service
Click "Create Model Service" and fill in the following information:
Service type: model service, machine learning service.
Basic configuration: service name, service version, service name description.
Model resources: select model, image resources.
Advanced setting: authentication enabled, enable service logs submission, configure COS storage address.
Resource configuration: select pay-as-you-go billing, CVM resource group.
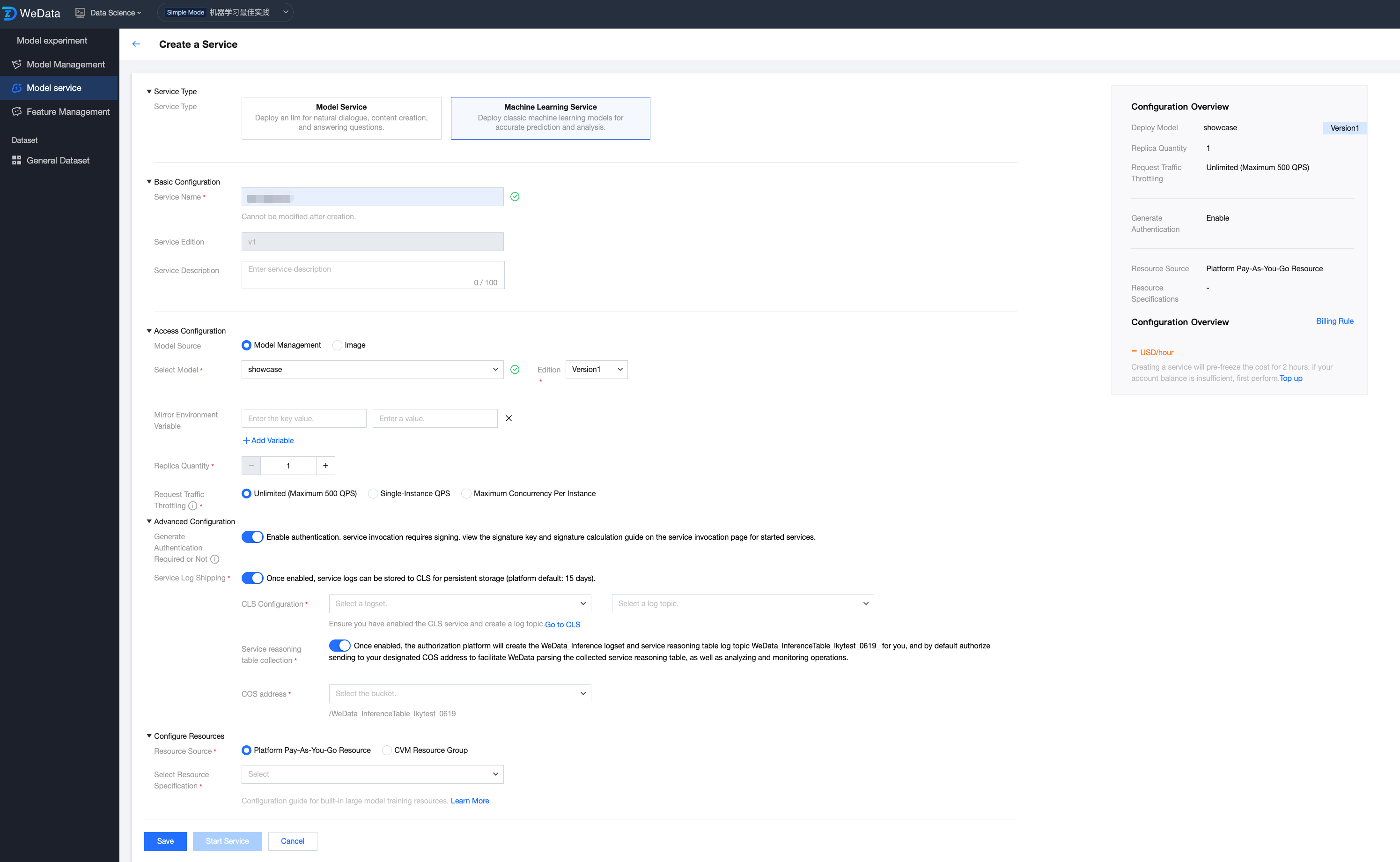
Model Source - Model Management
For models trained in a local development environment or WeData Notebook, users need to first register them to the model management module via the MLflow plug-in. After registration is successful, this model is converted to an enterprise-level standardized model asset, which can be called directly by the model service module and released as an online API with one click.
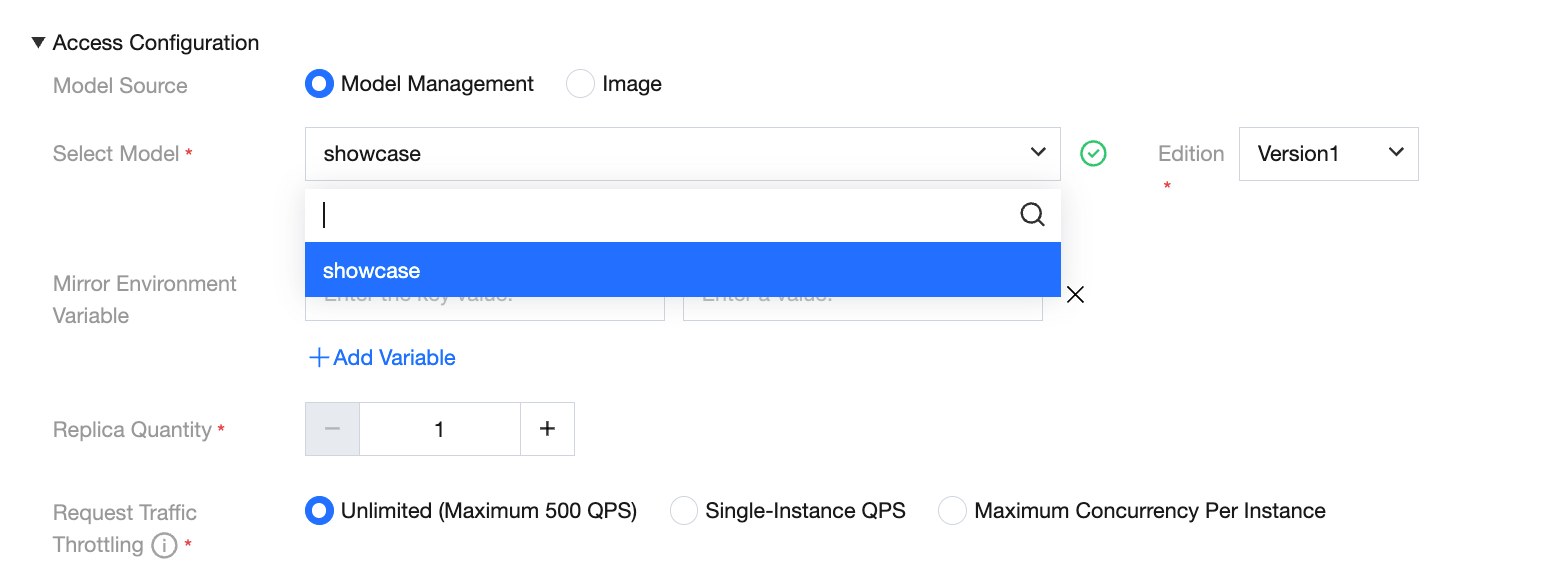
Model Source - Custom Image
WeData supports users to deploy model services via custom images, achieving high customization of the environment and deep integration of reasoning logic. For details, see Guide on Publishing Online Services Using Custom Images.
Core Values:
Deep custom environment: supports built-in non-standard machine learning frameworks or specified versions of system databases.
Preprocessing logic integration: allows direct encapsulation of "data stitching" or "feature online access" code in the mirror to implement an end-to-end closed development loop for "feature -> model -> reasoning".
Code mounting and flexible startup: Custom startup commands can overwrite the default behavior of the mirror and flexibly pull up different inference engines.
Import Method:
Tencent Container Registry (TCR) Enterprise Edition: suitable for in-house controlled mirror asset management, supports filtering by region and instance, down to namespace and repository URL.
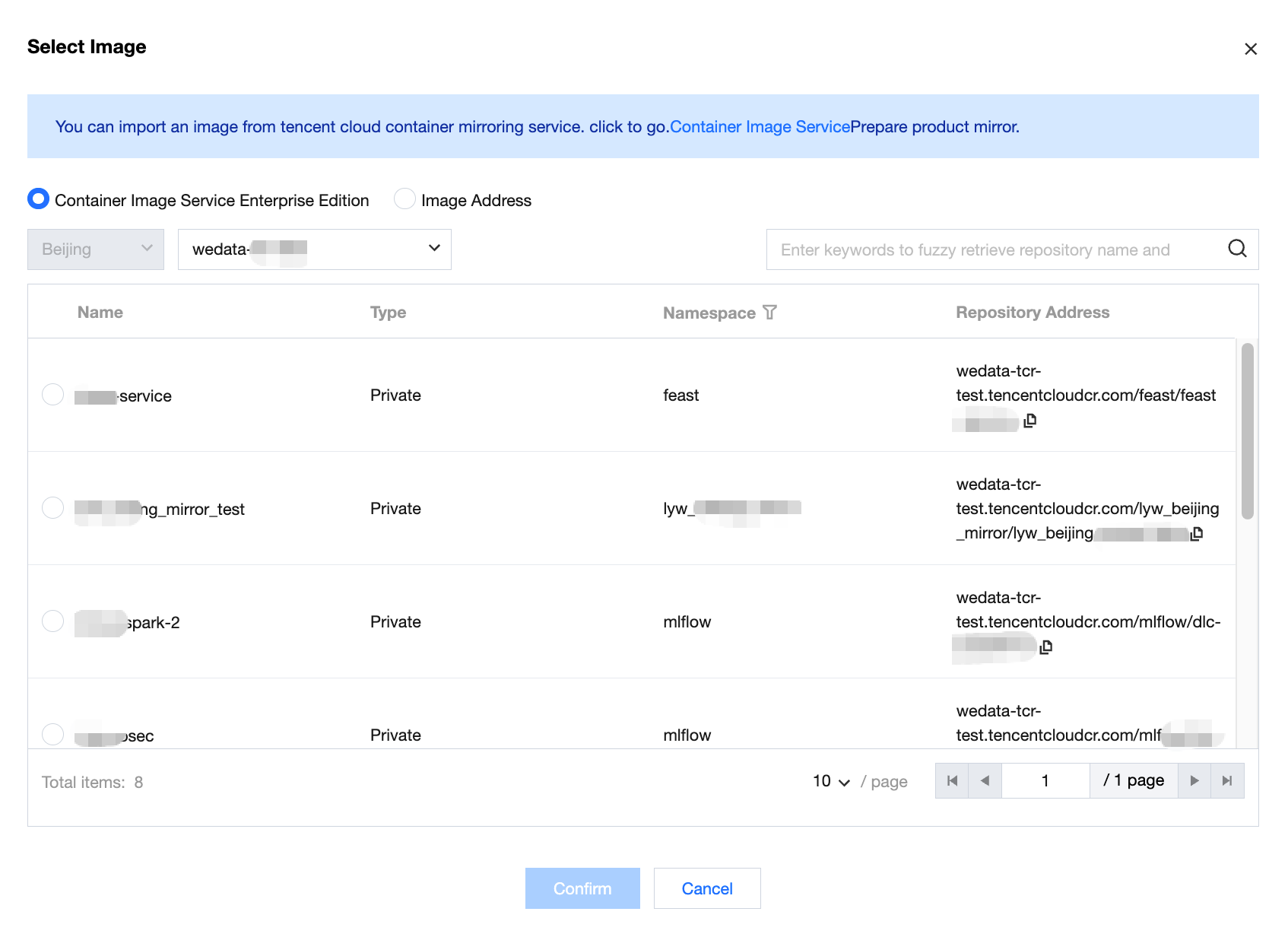
Image address: supports importing images from Tencent Cloud Container Registry.
Note:
Supports enabling authorization for private image repositories, which is off by default. After manually enabling, it supports input of the image repository username and password.
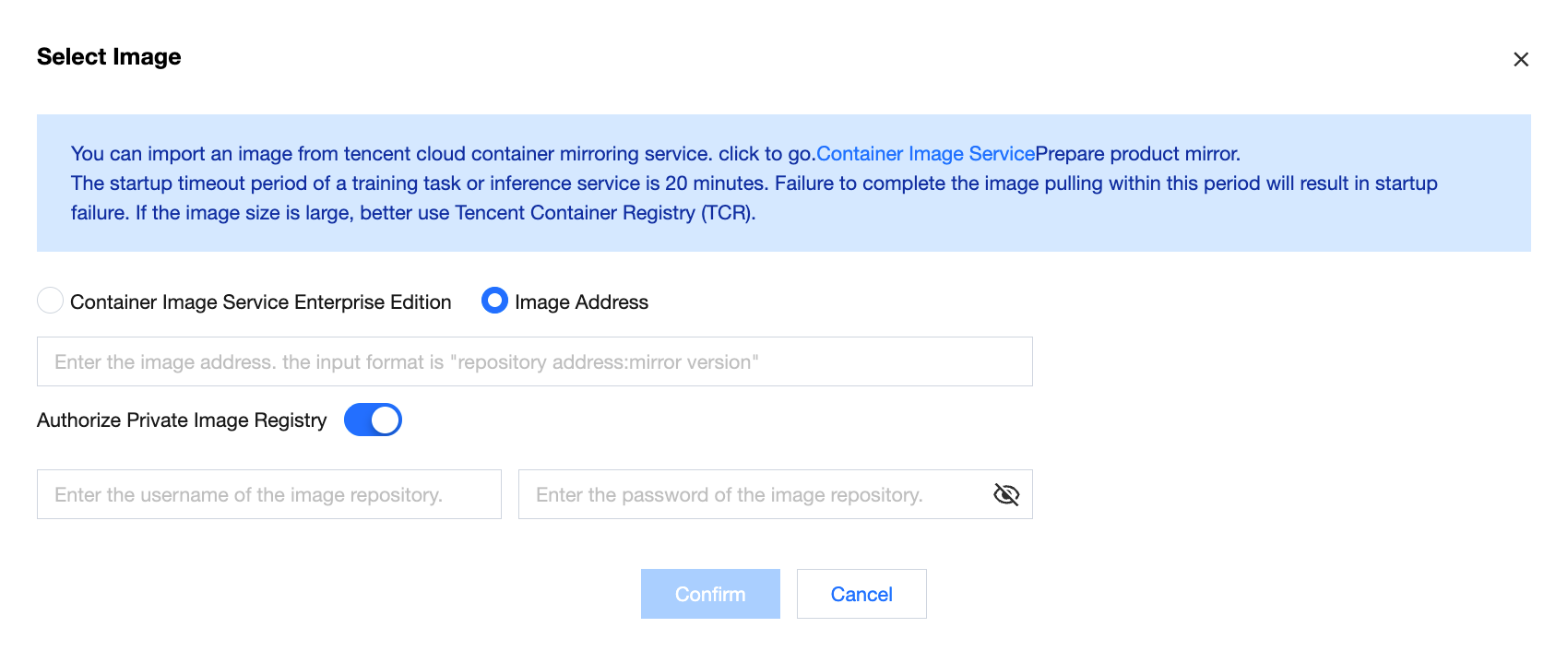
Enable Reasoning Table Collection (Supported Regions: Beijing, Shanghai, Chongqing, International-Singapore)
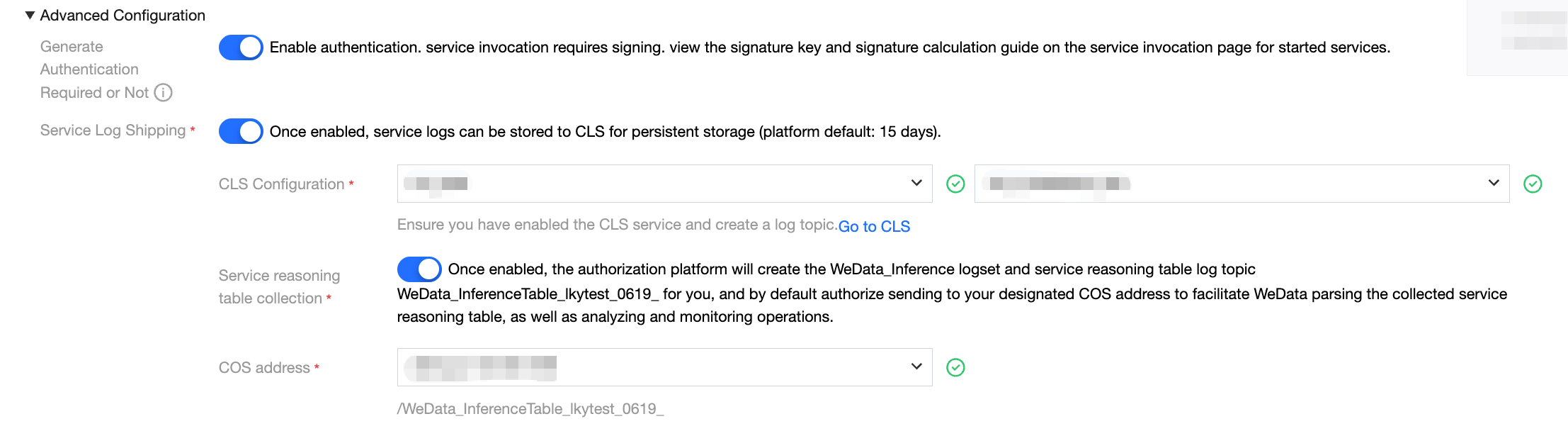
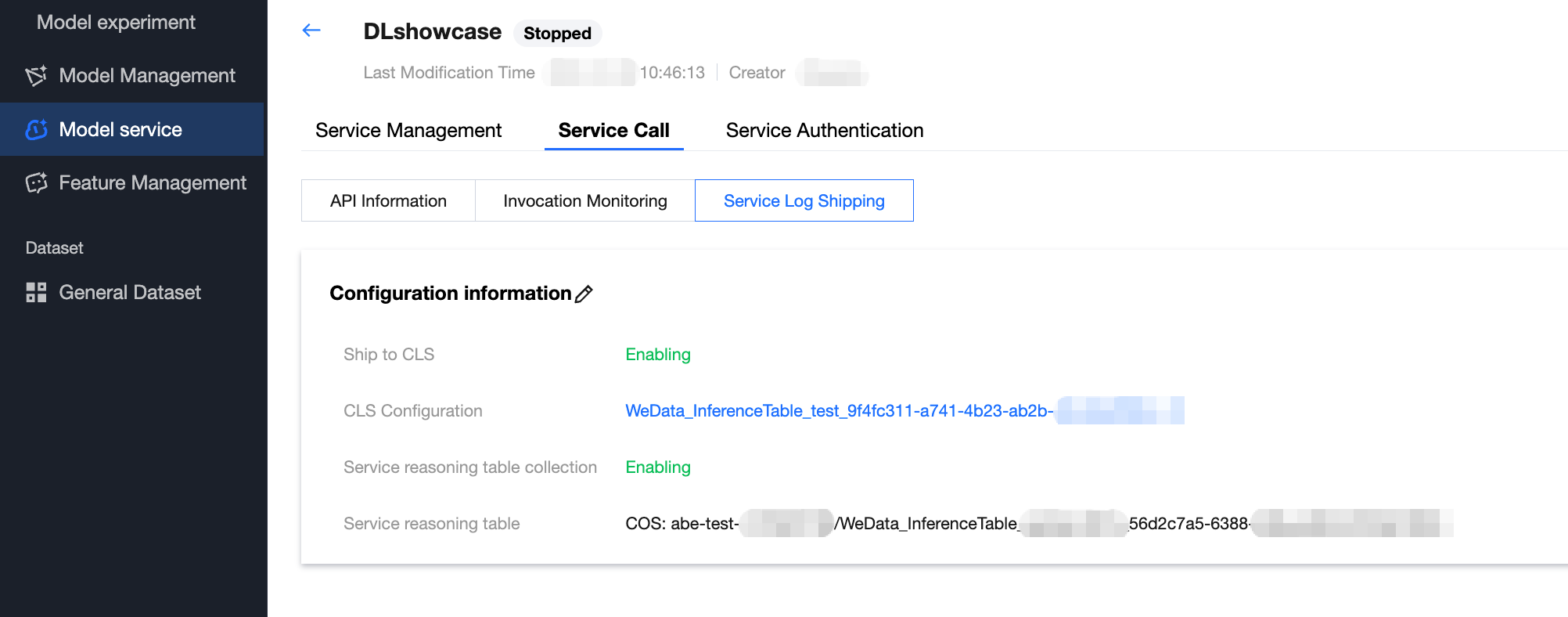
When creating a model service, complete log collection can be enabled through advanced configuration options.
Log delivery configuration: First, enable the log delivery feature and designate the destination log set and corresponding log topic. Once the configuration is complete, all request logs generated by the model service will be automatically delivered to the specified log topic.
Inference table log collection configuration: Enable the service inference table log collection option simultaneously and choose an appropriate COS bucket as the data storage destination. After enabling this feature, the system will automatically create a specialized log processing task responsible for filtering and structured processing of the collected full logs, then sending them to the pre-specified COS storage address.
Resource Source - Platform Pay-As-You-Go
The platform uses pay-as-you-go billing, providing a managed shared resource pool through WeData. Users do not need to worry about underlying server maintenance and can apply for calculation specifications on demand.
Operation Guide:
Select "platform pay-as-you-go resources" under "resource source".
In the "select resources" drop-down list, choose the appropriate specification based on model complexity.
Usage Scenario:
Development debugging: Quickly pull up service verification logic.
Tidal business: Severe traffic fluctuation with irregular intervals, hope to settle based on usage duration to optimize costs.
Lightweight reasoning: The model is small and does not require exclusive use of entire physical machine resources.

Resource Source - CVM Resource Group
CVM resource groups allow users to import purchased Tencent Cloud servers (CVM) into WeData for pool management, achieving physical-level isolation of computational resources.
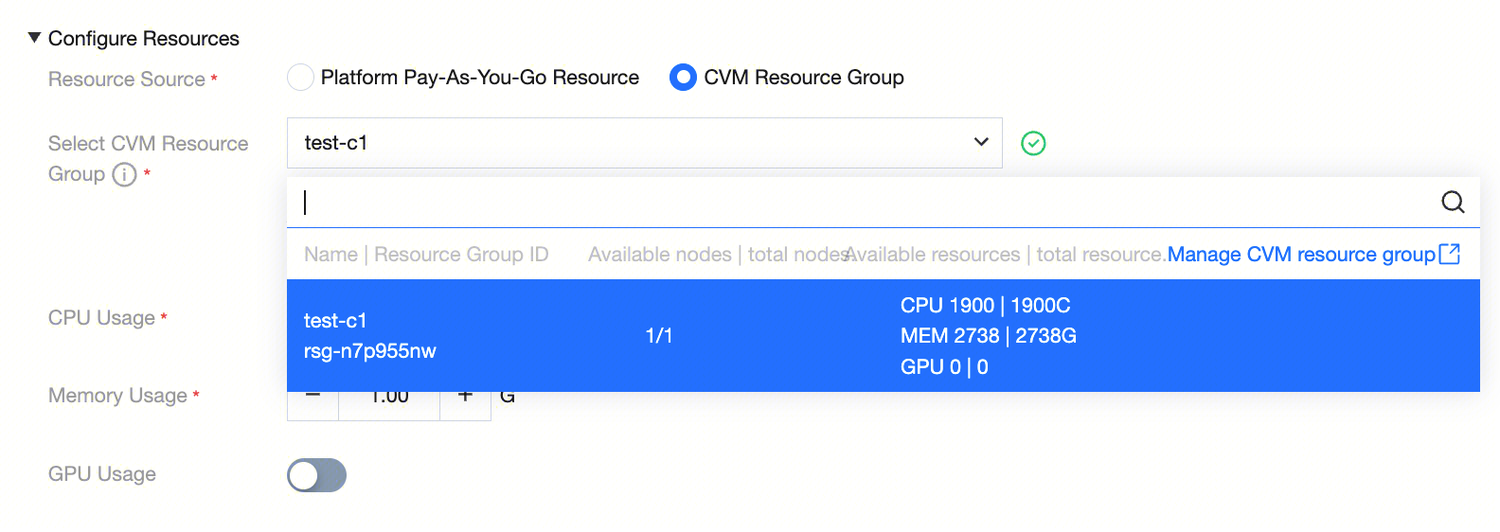
Operation Guide:
Check "CVM resource group" in "resource source".
Select the created and bound resource group from the dropdown list.
Note:
A resource group can only be associated with one service to ensure absolute exclusivity and stability of resources.
Use Cases:
Core production business: highly sensitive to delay, avoid noisy neighbor interference in shared environments.
High-concurrency reasoning: Maintain high QPS operation with long-term maintenance. Annual/monthly subscription CVM offers cost advantage.
GPU special project: Requires specifying specific model GPU computing cards (such as T4, A100) to perform high-performance reasoning.
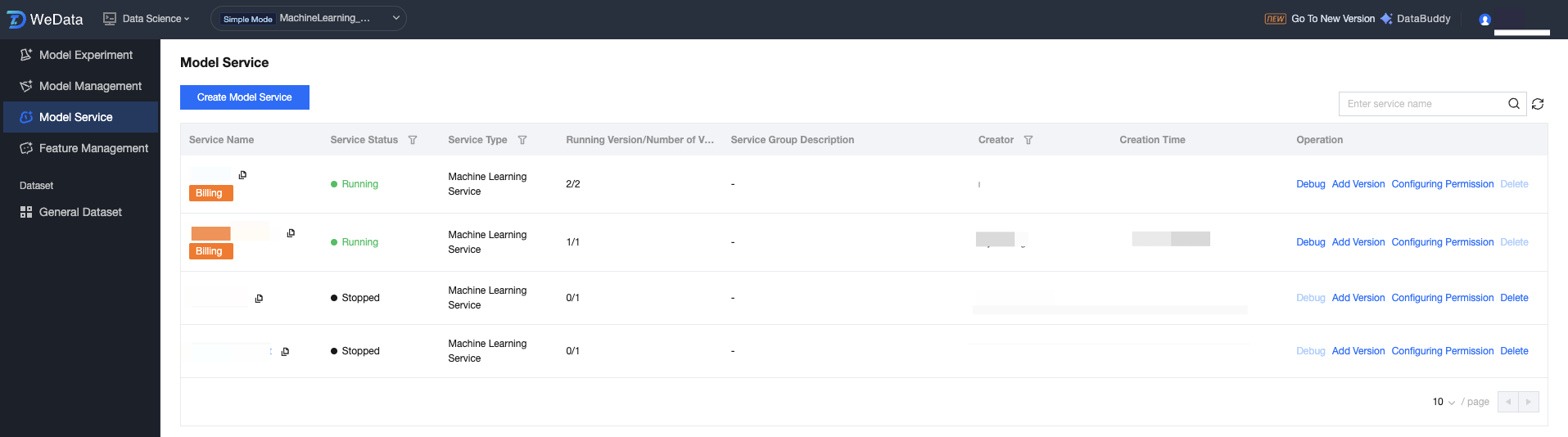
Model Service Group List
After the user creates a new service, all created model services are displayed in the list, including: service name, service status, description, creator, creation time, and operation method.
Field Classification | | Description(Optional) | Description |
Service Name | | The unique identifier name of the model service. | It is advisable to name according to business purpose. Click to go to the service details page. |
Service status | | Current lifecycle status of the service. | Common statuses include draft (starting), running, stopped. |
Service type | | Identify the application field of the service. | For example: machine learning service, model service. |
Running version/Number of versions | | Active service version/Total number of versions | Show the multi-version management of the service group, such as 0/1 means 1 version in total but not started. |
Service group description | | Description of the service group. | Facilitate team collaboration and quickly understand the purpose of the service group. |
Created by | | Create the user account for this service. | For auditing and permission trace. |
Creation time. | | The time when the service was first created. | Arranged in chronological order for quick search of recent services. |
Operation | Edit | Modify the basic configuration of the service group. | View service group details, support updating service group description and parameters such as resource configuration. |
| debug | Enter the online debugging page to test the API. | Provide a standardized REST API call address (service group must be in running state). |
| add version | Deploy a new model version under the same service. | Implement grayscale release or A/B testing scenarios for models, supporting up to two service versions. |
| Monitoring | View performance metrics. | Monitor QPS, number of concurrent requests, and CPU/MEM/GPU resource utilization rate. |
| Logs | View container logs at runtime. | Support by instance filtering and time range search for locating service group exception causes |
| Start | Control the running state of service groups. | Click Start to change the service group to running state. Stop releases service resources. |
| Deleted Object | Completely remove the model service asset and its associated resources. | The operation is irreversible. Confirm the service group has stopped running before deletion. |
Service Group Online Debugging
After deployment, the system provides standardized REST API information:
API Basic Info
Standard suffix address: The call address suffix for all services is fixed as /predict.
Request method: Use the POST method to call.
Data format: Use application/json format for both request and response.
Input Specification Description
During online testing and actual API calls, the input parameters must follow a specific JSON structure:
client_request_id: A unique identifier string for the request.
dataframe_split: A fixed key name used to encapsulate the feature data subject.
columns: A string list containing the field names of features defined during model training (its content is closely related to the specific model).
data: A nested list structure. The inner-layer list represents the eigenvalue of a single reasoning (must correspond to the sequence of columns). By placing multiple lists in parallel within the inner layer, multiple reasoning can be achieved per request.
Debugging the API Online Request Example
Taking the "Red Wine Quality Prediction" model as an example, the input JSON is as follows:
Single reasoning request body:
{"client_request_id": "123456","dataframe_split": {"columns": ["fixed_acidity", "volatile_acidity", "citric_acid", "residual_sugar", "chlorides", "free_sulfur_dioxide", "total_sulfur_dioxide", "density", "pH", "sulphates", "alcohol"],"data": [[1, 0.8, 0.7, 0.5, 1, 1.5, 0.8, 0.7, 0.5, 0.8, 0.5]]}}
Multiple reasoning (batch) request body: To predict two group data simultaneously, just append the eigenvalue list in the data array:
{"client_request_id": "123456","dataframe_split": {"columns": ["fixed_acidity", "volatile_acidity", "citric_acid", "residual_sugar", "chlorides", "free_sulfur_dioxide", "total_sulfur_dioxide", "density", "pH", "sulphates", "alcohol"],"data": [[1, 0.8, 0.7, 0.5, 1, 1.5, 0.8, 0.7, 0.5, 0.8, 0.5],[0.9, 0.7, 0.6, 0.4, 0.9, 1.2, 0.7, 0.6, 0.4, 0.7, 0.4]]}}
Calling the curl Command
Use a curl statement to enter, including the call address, authentication information (if any), content type, and input content as follows:
curl -X POST https://[service call address]/predict \\-H 'Authorization: [Authentication Token]' \\-H 'Content-Type: application/json' \\-d '{"client_request_id":"123456","dataframe_split":{"columns":["fixed_acidity","volatile_acidity","citric_acid","residual_sugar","chlorides","free_sulfur_dioxide","total_sulfur_dioxide","density","pH","sulphates","alcohol"],"data":[[1,0.8,0.7,0.5,1,1.5,0.8,0.7,0.5,0.8,0.5]]}}'
Call demo:
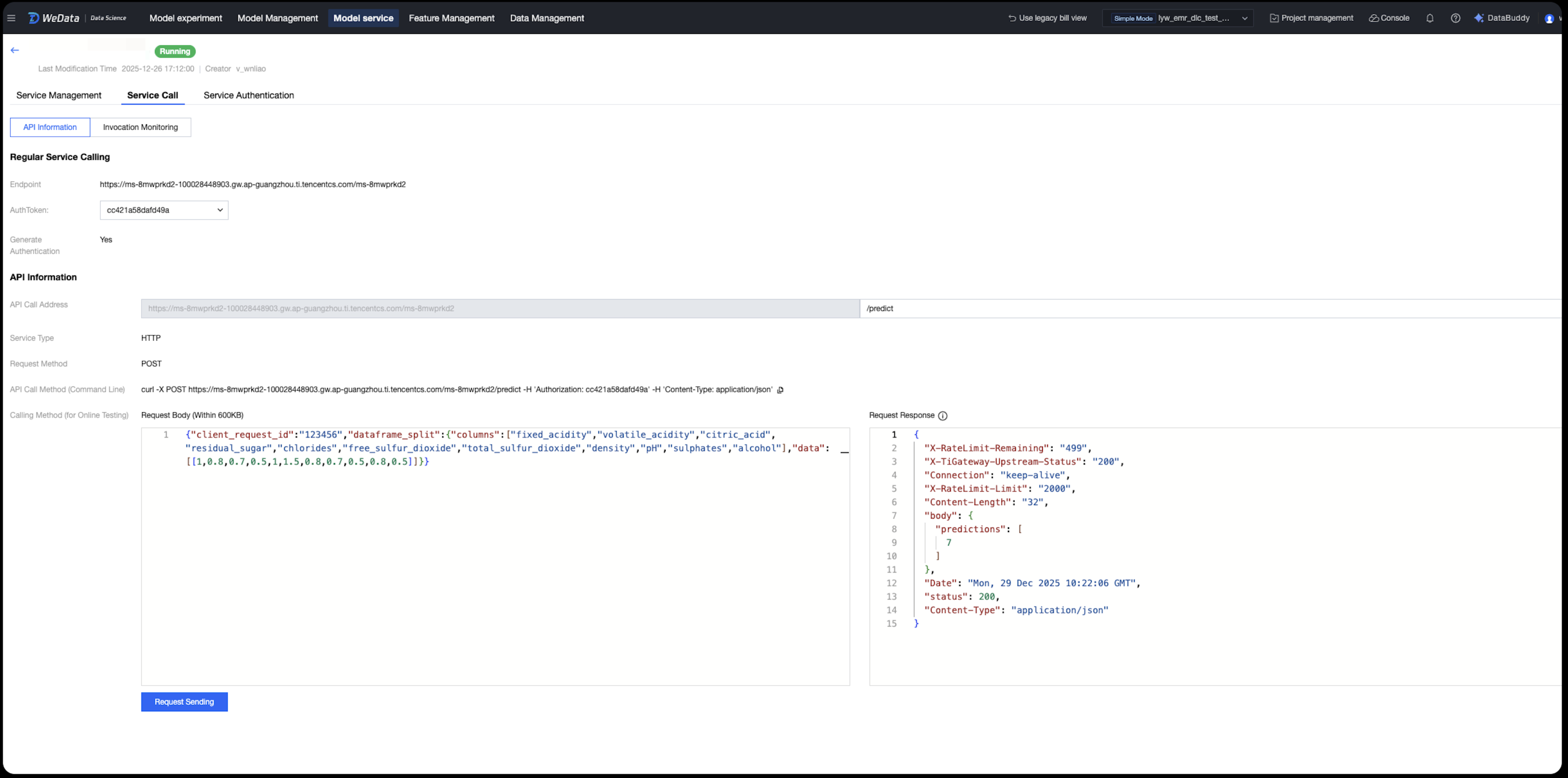
Service Group Performance Monitoring and Operations
The model service provides multi-dimensional monitoring views, supports viewing monitoring information (1 hour, 24 hours, 7 days, selected time period), and supports adjustments to time granularity (1 minute, 5 minutes):
Traffic monitoring: network traffic, QPS, QPS traffic throttling, number of concurrent requests.
Resource monitoring: CPU usage, memory usage, video memory utilization, GPU usage.
Instance management: instance count, number of running instances.
Service event: Records system events such as Pod scheduling and image pull, making it easy to troubleshoot startup failures.
Running log: Supports real-time refresh and search for log information output by model code.
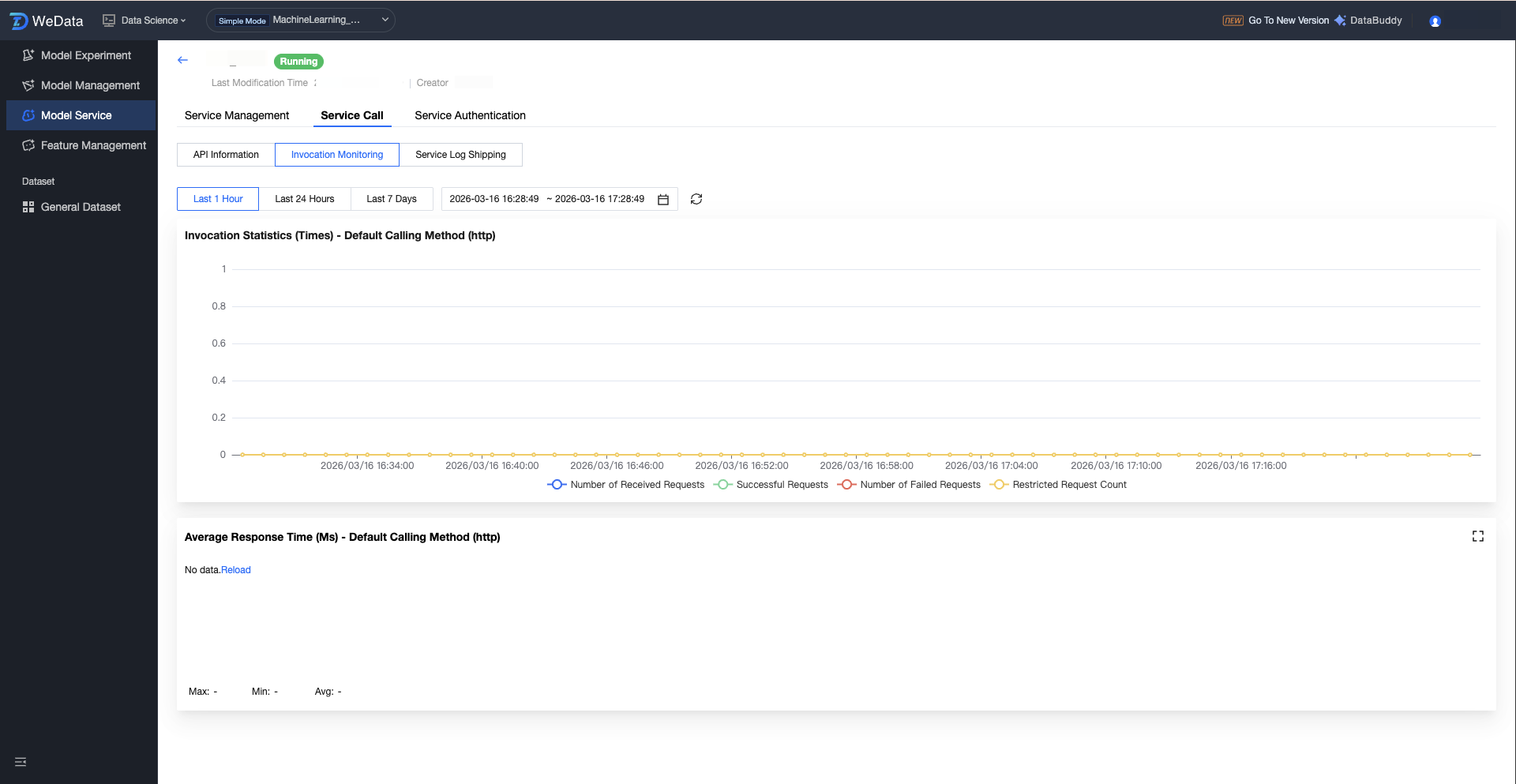
Service Group Enable Authentication Configuration
Service authentication is the core mechanism to ensure the security of model service calls. After enabling authentication, the system will perform identity verification on all requests entering the model service access point (Endpoint). Only requests carrying a valid key (API Key) will be allowed to access the model, thereby preventing illegal calls or malicious attacks on the model API. On the Version Details Page, switch to the "Service Authentication" tab to configure.
After enabling authentication: ALL REST API requests must contain the key in the Header.
After disabling authentication: The API is public, and any user with the endpoint can access it directly.
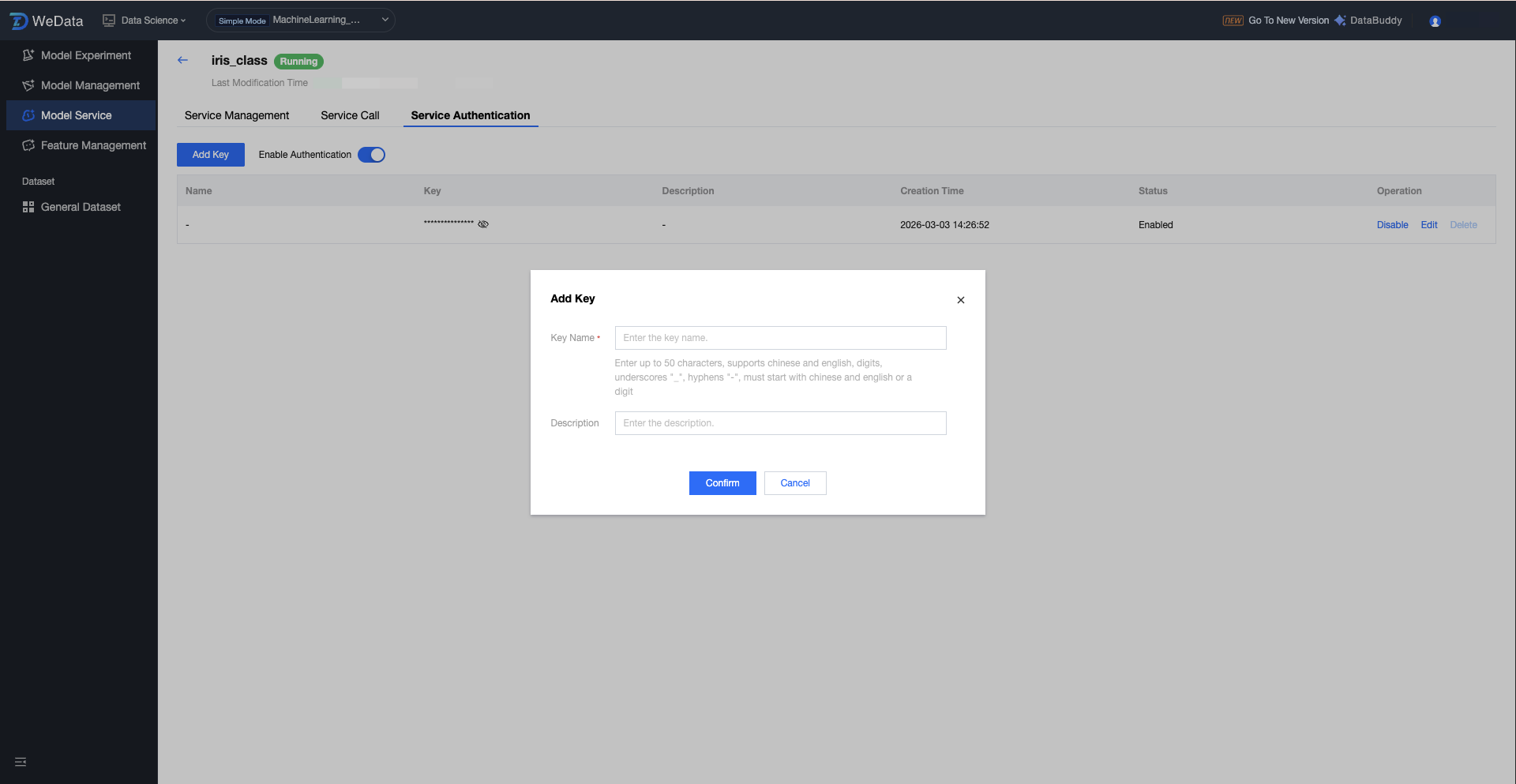
Service Version Operation Configuration
Click the service group name to enter the service version list, which shows all version information under the service group: ID, name, service version, status, description, model source, resource source, traffic allocation, creator, creation time, update time, and operations (update/monitor/logs/start/delete).
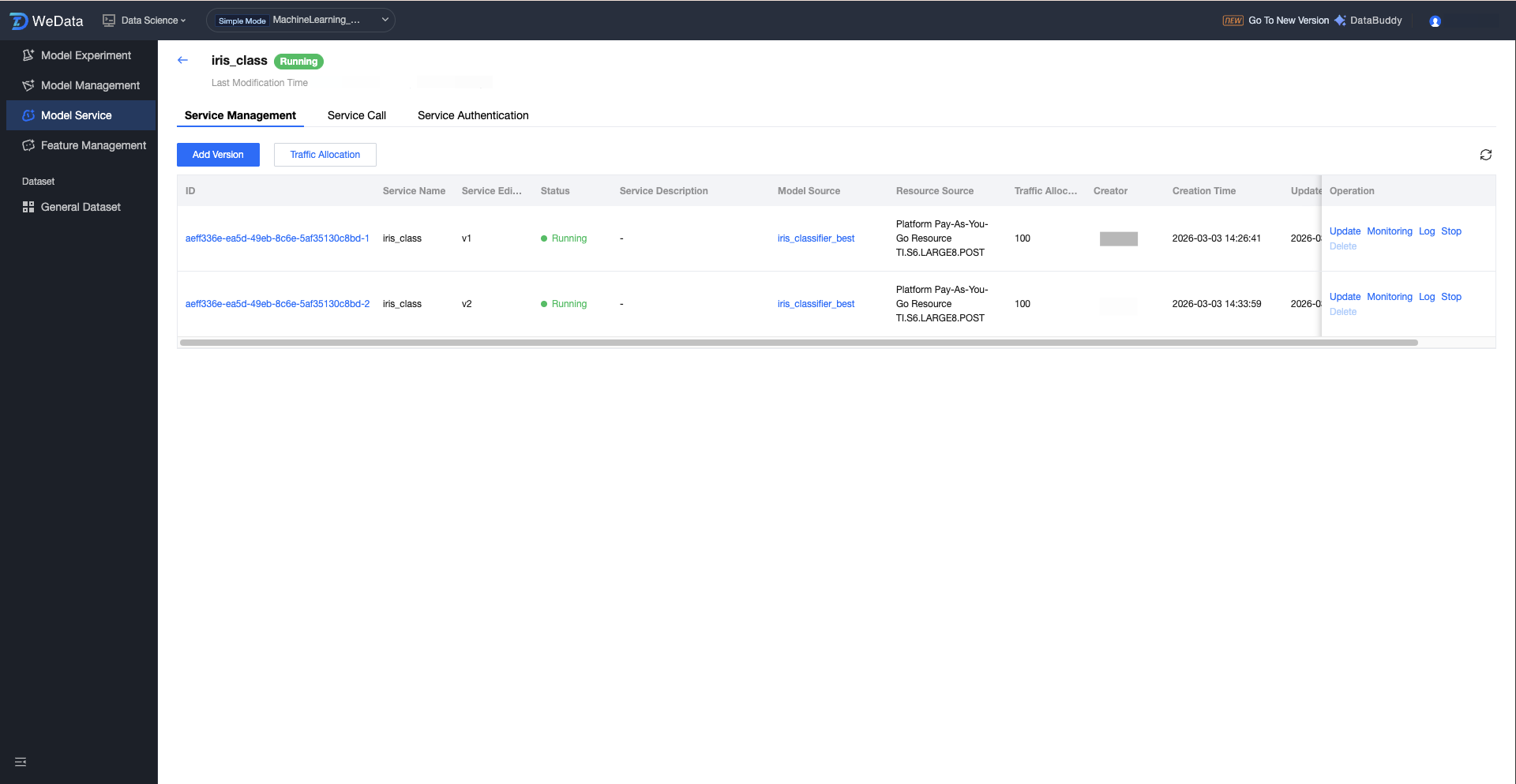
Field Classification | Description(Optional) | Description | |
ID | | Unique identifier of the service version instance. | A unique string automatically generated by the system for precise positioning and case-sensitive differentiation of service versions. |
Service Version Name | | The specific version flag of this service. | Facilitates multi-version management and grayscale release. |
State | | Current lifecycle status of the service version. | Show the version running state: draft/running/stopped. |
Description(Optional) | | Describe the purpose or feature of the service version. | Help understand service version differences in team collaboration. |
Model Source | | Publish the underlying model assets associated with the service. | Show which model name in the management module the service uses (for example, ml_wine_db_wine_model). |
Resource source | | The compute resource type consumed by this version during operation. | The resource is identified as coming from "platform pay-as-you-go resources" or "CVM resource group". |
Traffic Allocation | | The traffic weight received by this version during multi-version coexistence. | Displays the percentage of traffic assigned to this version in A/B testing or grayscale release scenarios. |
Created by | | Initialize the user account for this service version. | Record the creator of this version for permission management and CloudAudit. |
Creation time. | | The time when the service version was first created. | The system automatically records to track model iterations based on time. |
Operation | Updating | Modify the configuration message of the current service version. | View service version details, support updating version description and parameters such as resource configuration. |
| Monitoring | View real-time performance metrics of this version. | Visual monitoring view for QPS, call latency, CPU/GPU occupancy rate, etc. |
| Logs | Retrieve the running logs of the service instance. | Used to troubleshoot runtime issues such as model code errors and input/output exceptions. |
| Start/Stop | Manually control the running status of the service instance. | Click Start to change the service group to running state. Stop releases service resources. |
| Deleted Object | Permanently remove the service version. | The operation is irreversible. Confirm the version has stopped running before deletion. |
Service Version Detail
The Version Details Page provides a comprehensive view of the specified version model service, covering basic configuration, access parameters, and the running status of underlying container instances. Users can perform lifecycle management (start/stop), monitor in real time, and troubleshoot underlying issues on this page.
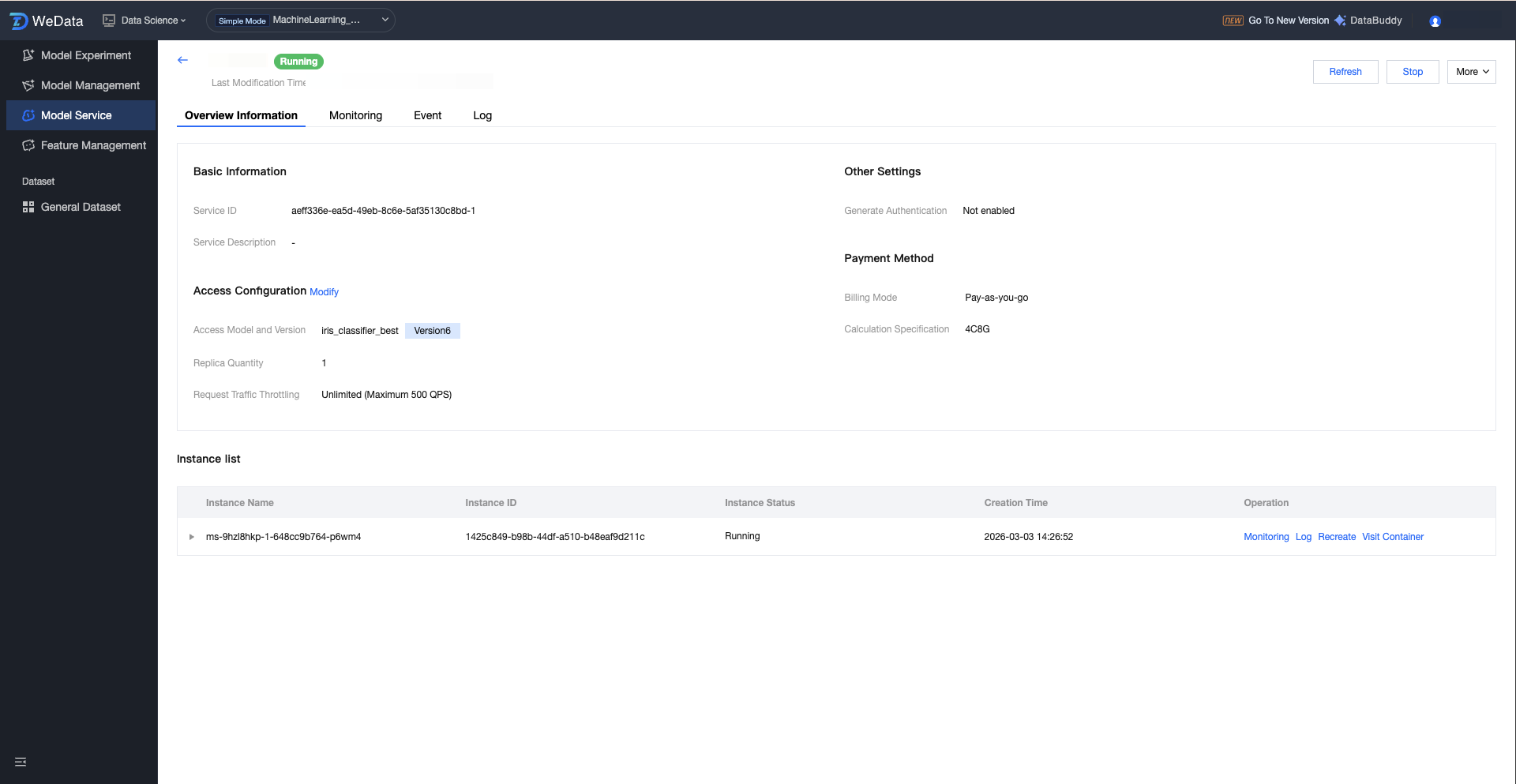
Service Version Details:
Field Name | Description(Optional) | Description |
Service Name | The specific version flag of this service. | Facilitates multi-version management and grayscale release. |
Service ID. | Unique identifier of the service version instance. | A unique string automatically generated by the system for precise positioning and case-sensitive differentiation of service versions. |
Service Description | Describe the purpose or feature of the service version. | Help understand service version differences in team collaboration. |
Billing Mode | Settlement method for resources | Pay-as-you-go shows the cost structure of model operation. |
Resource Specification | The computing power size is assigned to this version. | For example, 4C8G determines the model complexity and concurrency limit this version can handle. |
Access model and version | Associated model asset details. | Display the model name and specific MLflow version number (such as Version2) in model management. |
Number of Replicas | Total number of running container instances. | High availability of services is determined. Multi-replica can share traffic volume and provide disaster recovery protection. |
Request Traffic Throttling | Traffic protection threshold. | Show the QPS cap. If the threshold is reached, the system will trigger packet loss or queue protection. |
Generate Authentication | Service security control switch. | Display whether signature authentication (API Key) is enabled. If not enabled, the address can be accessed directly. |
Running instance details:
Field/Operation | Description(Optional) | Operations Guidance |
Instance name. | Identification of the underlying compute unit (Pod). | Assign a unique ID to a running service node. |
Instance State | Physical status of the instance. | Waiting indicates pulling images from registry or pending scheduling; Running means the service is ready. |
restart attempt | How often the container exits abnormally. | Core troubleshooting metrics. If the count keeps increasing, the model may contain memory leaks or code errors. |
Monitoring | Instance-level monitoring entry. | View individual instance $CPU/MEM$ consumption and check whether resource skew exists. |
Logs | Container standard output stream. | View model load logic and predict error stack with the most direct method. |
Restart | Force reboot an instance. | An urgent method to quickly restore services when an instance becomes dead or responds extremely slow. |
Access a container | Interactive terminal. | Allow entering the container to run commands, check file path, permission, or environment dependency. |
Service Version Monitoring
The monitoring page provides real-time perceptual ability for the service running status of the model, tracking service performance from three dimensions: business traffic, system resources, and underlying instances. Through visual charts, it helps algorithm engineers and O&M personnel quickly locate performance bottlenecks, verify scaling effects, and assess the response quality of the model.
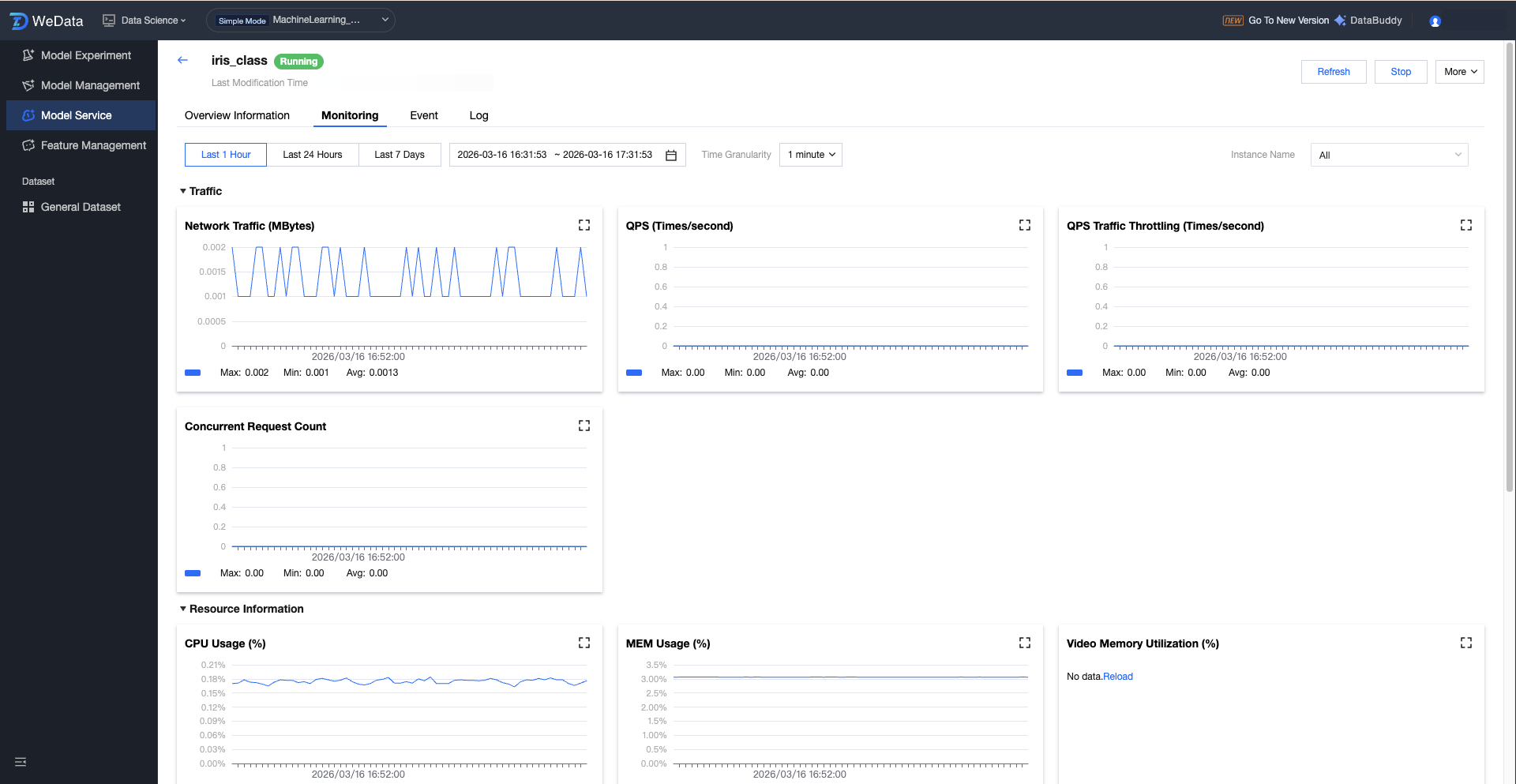
Metric Category | Core Metric Items | Business Value Description |
Traffic information | Network traffic, QPS, QPS throttling count, number of concurrent requests | Assess API call pressure. Concurrency request count and throttling count are key signals to assess whether to increase replica quantity. |
Resource information | CPU usage, MEM usage, video memory utilization, GPU usage | Monitor underlying hardware load, including video memory utilization and GPU usage. |
Instance information | Total number of instances Number of running instances | Intuitively display service availability and confirm whether the actual number of containers running conforms to the expected configuration. |
Service Version Events and Logs
Service Operation Event:
Support viewing operational service events, including first occurrence time, last event, level, resource type, resource name, detailed description, occurrence count, etc.
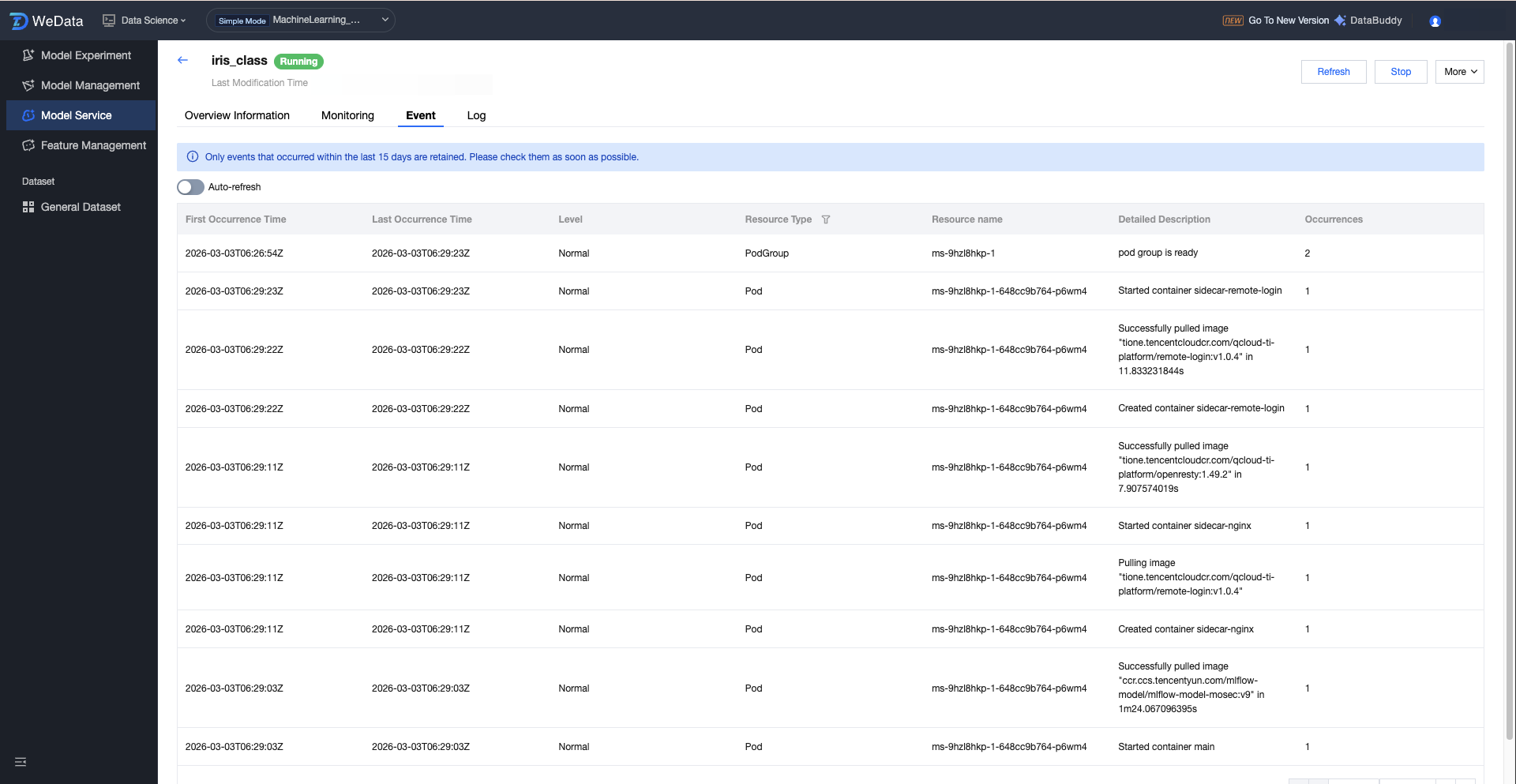
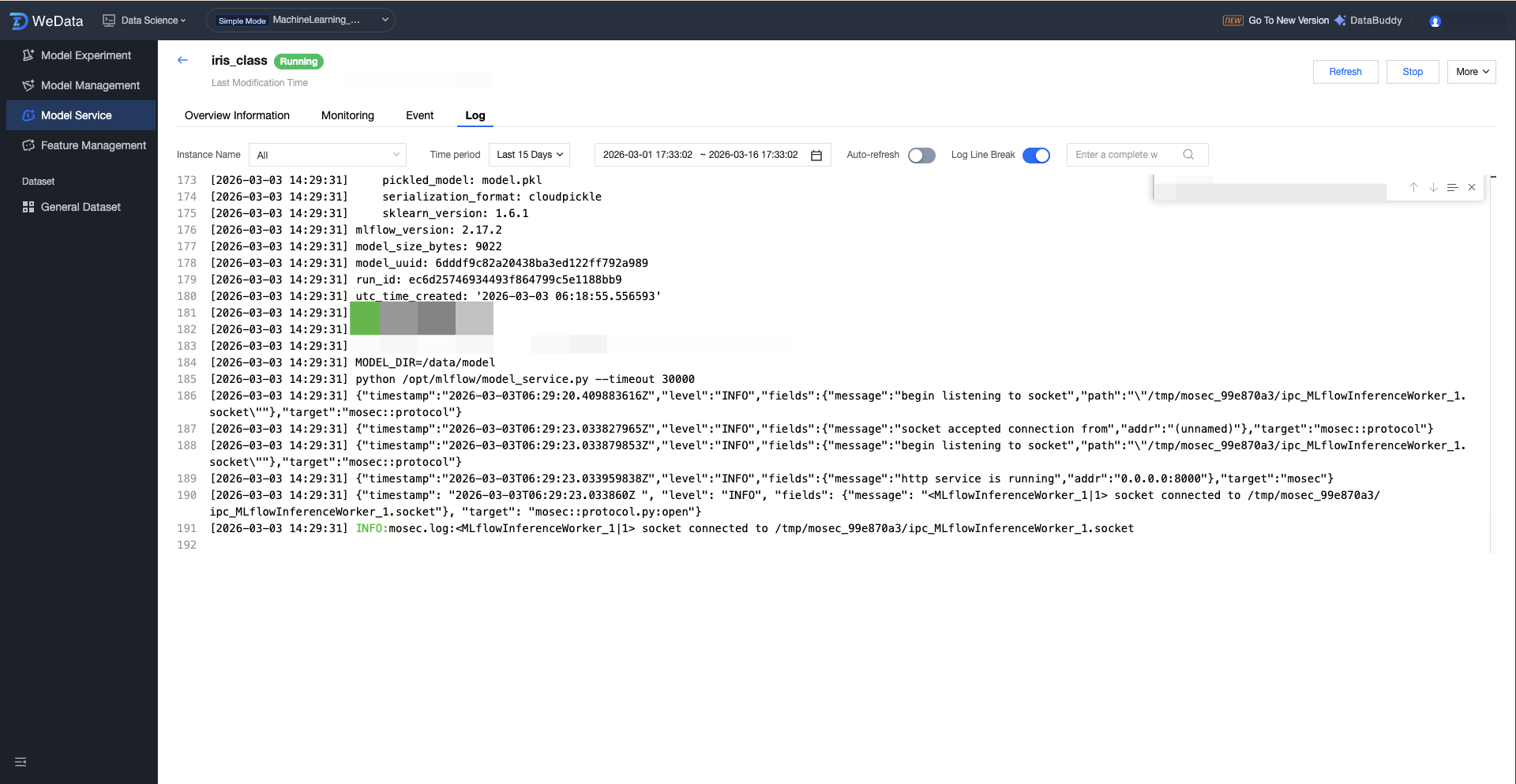
Service Operation Logs:
Supports viewing service logs, filtering by instance and time range (1 hour, 24 hours, 7 days, 15 days, selected time), and features like automatically refresh, search, and log line break.
Appendix
Model Monitoring Best Practice
What Is a Reasoning Table
The inference table is an important tool for machine learning model quality monitoring, enabling the Ops team to monitor the running state and performance of online models in real time. As shown below, in a typical workflow, when the monitoring function is enabled in the model service, the system will automatically capture and record key information during the model inference process, including feature data, model output, inference latency, and other key metrics.
The recorded data is written into the inference table in real time, providing a data foundation for subsequent quality analysis. By collaborating with the data quality monitoring module, multidimensional quality checks can be performed on the data in the table, including data distribution change detection, model prediction accuracy monitoring, and abnormal value identification. When the monitoring system detects a downward trend or anomalies in model quality, it promptly triggers an alarm mechanism to remind related personnel to take appropriate optimization measures, such as adjusting model parameters, updating the training dataset, or retraining the model.
The entire process forms a closed-loop from data acquisition, quality monitoring to problem response, ensuring online models can continuously and stably provide high-quality service, while offering data support for continuous optimization and iteration of models.
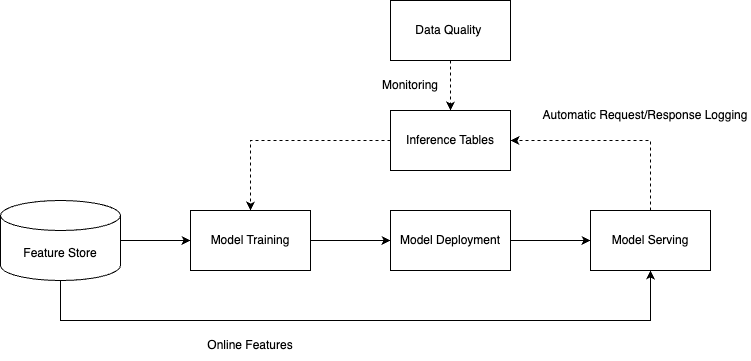
Reasoning Table Monitoring Process
The following diagram shows the complete technical flow of inference table data monitoring. The entire process is divided into three core stages: data acquisition, data processing, and quality monitoring.
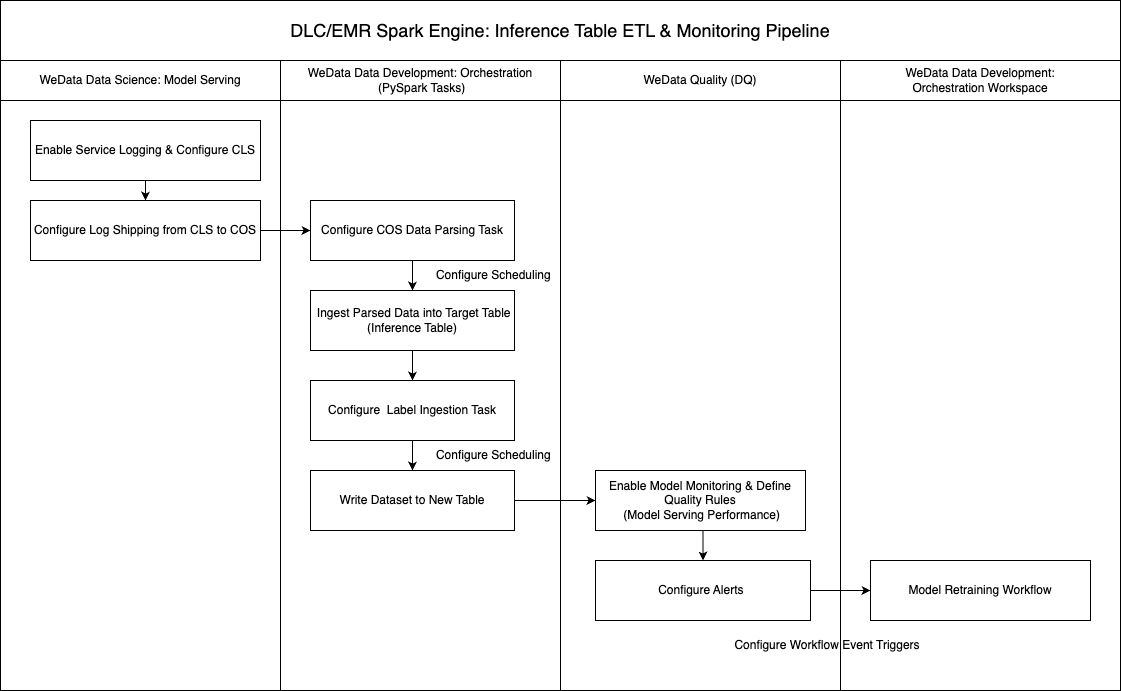
Data acquisition stage: During model service deployment, the log collection feature and log delivery mechanism must be enabled. The system will automatically collect various log information generated during model inference and securely transmit the raw log data to COS via the configured delivery strategy.
Data processing stage: After log data is successfully delivered to COS, you can configure PySpark data processing tasks in the orchestration space of WeData offline development. By writing appropriate data processing logic, the original inference logs are scrubbed, switched, and structured to extract key metrics such as model input features, prediction results, response time, and error information. The standardized inference data is then written to a specialized inference table.
Quality monitoring stage: Based on structured data in the inference table, configure service quality monitoring rules in the WeData data quality module, setting multi-level monitoring metrics and alarm thresholds. If model performance anomalies, data drift, or service quality degradation are detected, the system will automatically trigger alarm notifications. Furthermore, based on the severity and type of alarm events, it can automatically trigger corresponding model retraining workflows, achieving an intelligent Ops closed-loop from problem discovery to automatic fixes.
Data Acquisition
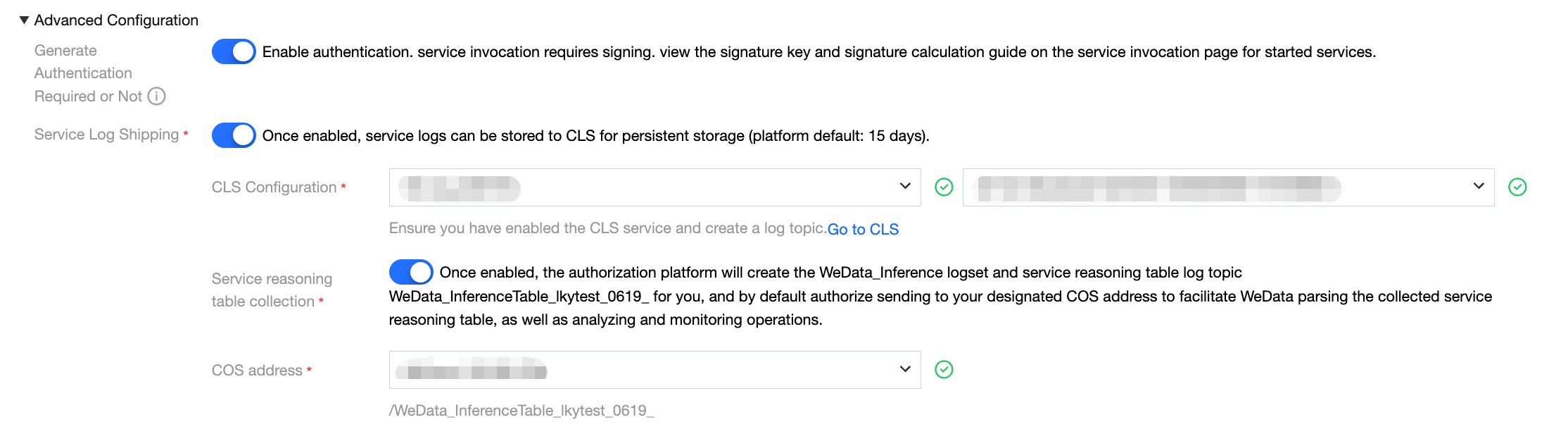
When creating a model service, complete log collection can be enabled through advanced configuration options.
Service log delivery configuration: First, enable the service log delivery feature and designate the target logset and corresponding log topic. Once the configuration is complete, all request logs generated by the model service will be automatically delivered to the designated log topic.
Inference table log collection configuration: Enable the service inference table log collection option simultaneously and choose an appropriate COS bucket as the data storage destination. After enabling this feature, the system will automatically create a specialized log processing task responsible for filtering and structured processing of collected full logs, then sending them to the pre-specified COS storage address.
Data Processing
Processing COS Log Data for Reasoning
The purpose of this step is to parse the original model input and output information, convert unstructured log data into standardized feature data and prediction results, and write the processed data to a specialized inference table, providing a structured data basis for subsequent data analysis and tag association work.
Technical implementation plan:
Stream processing architecture: Recommended to use Spark Streaming for real-time streaming data processing. This method can continuously monitor new log files in the COS bucket and perform real-time data parsing and switching to ensure data timeliness.
Incremental processing mechanism: Configure a Checkpoint mechanism during data writing to record processing progress and status info. By setting a scheduled run policy, the system can automatically identify and process incremental data, avoid duplicate processing of existing data, and ensure data processing integrity and consistency.
Fault tolerance assurance: The Checkpoint mechanism also provides failure recovery capability. When a processing task is interrupted due to exceptions, it can resume execution from the latest checkpoint to ensure data processing reliability.
Implementation code can be based on different engines as follows:
DLC Spark Engine
This example demonstrates how to use Spark Streaming to read the reasoning table in COS storage in streaming mode, process key fields (feature, predicted value), and write to the DLC internal table.## PrerequisiteEnsure the COS access credential associated with the DLC engine has permission to access the reasoning table storage bucket.2. Underwrite the model service has enabled reasoning table monitoring.## How to runAfter modifying parameters in the tutorial, select DLC PySpark task from offline development > orchestration space to run.## Scheduled jobAfter modifying the corresponding parameters and completing debugging, you can configure the task for scheduled execution to continuously process reasoning table data. You can also associate the processed reasoning table data with tag data to generate a reasoning table containing tag data, and configure machine learning-related monitoring metrics for the reasoning table in the data quality module to continuously monitor model quality.""" from pyspark.sql import DataFrame, SparkSessionfrom pyspark.sql import functions as Ffrom pyspark.sql import types as Tfrom typing import Union, List, Dict, Optional, Sequence, Anyfrom pyspark.sql.streaming import StreamingQueryfrom pyspark.sql.types import (StructType, StructField, StringType, LongType, DoubleType,ArrayType, TimestampType)"""### Define original reasoning table data formatThe original reasoning table data is stored in the user-specified COS bucket in JSON format with a specific structure. request_schema represents the input request structure, and response_schema represents the model output structure. Adjust according to the actual input and output format of the model service. Other fields are part of the fixed structure of the reasoning table and generally not needed to adjust."""request_schema = StructType([StructField("client_request_id", StringType(), True),StructField("dataframe_split", StructType([StructField("columns", ArrayType(StringType()), True),StructField("data", ArrayType(ArrayType(DoubleType())), True),]), True),])request_metadata_schema = StructType([StructField("model_name", StringType(), True),StructField("model_version", StringType(), True),StructField("service_id", StringType(), True),])response_schema = StructType([StructField("predictions", ArrayType(LongType()), True),])schema = StructType([StructField("client_request_id", StringType(), True),StructField("execution_duration_ms", LongType(), True),StructField("request", request_schema, True),StructField("request_date", StringType(), True),StructField("request_metadata", request_metadata_schema, True),StructField("request_time", TimestampType(), True),StructField("requester", StringType(), True),StructField("response", response_schema, True),StructField("sampling_fraction", DoubleType(), True),StructField("status_code", LongType(), True),StructField("wedata_request_id", StringType(), True),])# data source parametersSOURCE_PATH = "cosn://bucket path/" # storage path of the reasoning tableCHECKPOINT_PATH = "cosn://bucket path/checkpoint" # storage path of check_point, recommended to match the reasoning table storage path# result table parametersUNPACKED_TABLE_NAME = "DataLakeCatalog.test.unpacked_test_inference_table" # Result table name, need to be a three-segment table name in DLC: <data catalog>.<database>.<data table>MODEL_ID_COL = "model_id" # model ID column in the result table, used for identification modelEXAMPLE_ID_COL = "example_id" # unique ID column for result table entries, used for update operationsPREDICTION_COL = "prediction" # column name for predicted valueFEATURE_COLUMNS = ["sepal_length", "sepal_width", "petal_length", "petal_width"] # feature column names, adjust based on model featuresdef process_requests(requests_raw: DataFrame) -> DataFrame:"""Expand request features into separate columns and pair them with prediction resultsArgs:requests_raw: pending dataReturn:processed data"""filter successful requestsrequests_success = requests_raw.filter(F.col("status_code") == 200).drop("status_code")# Generate model flagrequests_identified = requests_success \\.withColumn(MODEL_ID_COL, F.concat(F.col("request_metadata").getItem("model_name"), F.lit("_"), F.col("request_metadata").getItem("model_version"))) \\.drop("request_metadata")# Unfold feature column and prediction result# 1. Get feature column names, feature data, predicted valuerequests_with_features = (requests_identified.withColumn("feature_columns", F.col("request.dataframe_split.columns")).withColumn("feature_data", F.col("request.dataframe_split.data")).withColumn("predictions", F.col("response.predictions")))# 2. Pair each data row with the corresponding prediction resultrequests_exploded = (requests_with_features.withColumn("feature_prediction_pairs",F.arrays_zip(F.col("feature_data"), F.col("predictions"))).withColumn("feature_prediction_pairs",F.explode(F.col("feature_prediction_pairs"))).withColumn("feature_row", F.col("feature_prediction_pairs.feature_data")).withColumn(PREDICTION_COL, F.col("feature_prediction_pairs.predictions")))# 3. dynamically create feature columnrequests_with_feature_cols = requests_explodedfor i, col_name in enumerate(FEATURE_COLUMNS):requests_with_feature_cols = (requests_with_feature_cols.withColumn(col_name, F.col("feature_row")[i]))# 4. Add record unique ID, making it easy to perform upsert operationrequests_with_example_id = requests_with_feature_cols \\.withColumn(EXAMPLE_ID_COL, F.expr("uuid()"))# 5. Clean up temporary columnsrequests_processed = (requests_with_example_id.drop("feature_columns", "feature_data", "feature_prediction_pairs", "feature_row", "predictions", "request", "response"))return requests_processeddef create_table(spark: SparkSession,table_name: str,df: Optional[DataFrame] = None,partition_expr: str = None,description: Optional[str] = None,):"""Create a dlc internal tableArgs:spark: SparkSession instancetable_name: full table name (format: data catalog.database.data table)df: initial data (used for infer schema)partition_expr: partition column (optimized storage query)description: table descriptionRaises:ValueError: throw create exception"""# Infer table schematable_schema = df.schema# Build column definitioncolumns_ddl = []for field in table_schema.fields:data_type = field.dataType.simpleString().upper()col_def = f"`{field.name}` {data_type}"if not field.nullable:col_def += " NOT NULL"columns_ddl.append(col_def)# Build table creation statementddl = f"""CREATE TABLE IF NOT EXISTS {table_name} ({', '.join(columns_ddl)})USING icebergPARTITIONED BY ({partition_expr})TBLPROPERTIES ('comment'= '{description or ''}','format-version'= '2','write.metadata.previous-versions-max'= '100','write.metadata.delete-after-commit.enabled'= 'true','smart-optimizer.inherit' = 'none','smart-optimizer.written.enable' = 'enable')"""# Print sqlprint(f"create table ddl: {ddl}\\n")# execute DDLtry:spark.sql(ddl)except Exception as e:raise ValueError(f"Failed to create table: {str(e)}") from eprint(f"create table {table_name} done")spark = SparkSession.builder.appName("Operate DB Example").getOrCreate()# Build a streaming DataFramedf = (spark.readStream.format("json").schema(schema).load(SOURCE_PATH))# Process streaming dataunpacked_df = process_requests(df)# Create a result table with request_time as partition field, using daily partitioning, which can be adjusted by service request volumecreate_table(spark, UNPACKED_TABLE_NAME, unpacked_df, 'days(request_time)')# Write dataquery = unpacked_df.writeStream \\.format("parquet") \\.outputMode("append") \\.option("checkpointLocation", CHECKPOINT_PATH) \\.trigger(once=True) \\.toTable(UNPACKED_TABLE_NAME)query.awaitTermination()spark.sql(f"select * from {UNPACKED_TABLE_NAME}").show(10)
EMR Spark Engine
"""This example demonstrates how to use Spark Streaming to read the reasoning table from COS storage in streaming mode, process key fields (feature, predicted value), and write to the EMR Hive table.## PrerequisiteEnsure the COS access credential associated with the DLC engine has permission to access the reasoning table storage bucket.2. Underwrite the model service has enabled reasoning table monitoring.## How to runAfter modifying parameters in the tutorial, select EMR PySpark task from offline development > orchestration space to run.## Scheduled jobAfter modifying the corresponding parameters and completing debugging, you can configure the task for scheduled execution to continuously process reasoning table data. You can also associate the processed reasoning table data with tag data to generate a reasoning table containing tag data, and configure machine learning-related monitoring metrics for the reasoning table in the data quality module to continuously monitor model quality."""from pyspark.sql import DataFrame, SparkSessionfrom pyspark.sql import functions as Ffrom pyspark.sql import types as Tfrom typing import Union, List, Dict, Optional, Sequence, Anyfrom pyspark.sql.streaming import StreamingQueryfrom pyspark.sql.types import (StructType, StructField, StringType, LongType, DoubleType,ArrayType, TimestampType)"""### Define original reasoning table data formatThe original reasoning table data is stored in the user-specified COS bucket in JSON format with a specific structure. request_schema represents the input request structure, and response_schema represents the model output structure. Adjust according to the actual input and output format of the model service. Other fields are part of the fixed structure of the reasoning table and generally not needed to adjust."""request_schema = StructType([StructField("client_request_id", StringType(), True),StructField("dataframe_split", StructType([StructField("columns", ArrayType(StringType()), True),StructField("data", ArrayType(ArrayType(DoubleType())), True),]), True),])request_metadata_schema = StructType([StructField("model_name", StringType(), True),StructField("model_version", StringType(), True),StructField("service_id", StringType(), True),])response_schema = StructType([StructField("predictions", ArrayType(LongType()), True),])schema = StructType([StructField("client_request_id", StringType(), True),StructField("execution_duration_ms", LongType(), True),StructField("request", request_schema, True),StructField("request_date", StringType(), True),StructField("request_metadata", request_metadata_schema, True),StructField("request_time", TimestampType(), True),StructField("requester", StringType(), True),StructField("response", response_schema, True),StructField("sampling_fraction", DoubleType(), True),StructField("status_code", LongType(), True),StructField("wedata_request_id", StringType(), True),])# data source parametersSOURCE_PATH = "cosn://bucket path/" # storage path of the reasoning tableCHECKPOINT_PATH = "cosn://bucket path/checkpoint" # storage path of check_point, recommended to match the reasoning table storage path# result table parametersUNPACKED_TABLE_NAME = "testdb.UNPACKED_test_inference_TABLE" # Result TABLE NAME, need to be a hive TABLE NAME: <database>.<data TABLE>MODEL_ID_COL = "model_id" # model ID column in the result table, used for identification modelEXAMPLE_ID_COL = "example_id" # unique ID column for result table entries, used for update operationsPREDICTION_COL = "prediction" # column name for predicted valueFEATURE_COLUMNS = ["sepal_length", "sepal_width", "petal_length", "petal_width"] # feature column names, adjust based on model featuresdef process_requests(requests_raw: DataFrame) -> DataFrame:"""Expand request features into separate columns and pair them with prediction resultsArgs:requests_raw: pending dataReturn:processed data"""filter successful requestsrequests_success = requests_raw.filter(F.col("status_code") == 200).drop("status_code")# Generate model flagrequests_identified = requests_success \\.withColumn(MODEL_ID_COL, F.concat(F.col("request_metadata").getItem("model_name"), F.lit("_"), F.col("request_metadata").getItem("model_version"))) \\.drop("request_metadata")# Unfold feature column and prediction result# 1. Get feature column names, feature data, predicted valuerequests_with_features = (requests_identified.withColumn("feature_columns", F.col("request.dataframe_split.columns")).withColumn("feature_data", F.col("request.dataframe_split.data")).withColumn("predictions", F.col("response.predictions")))# 2. Pair each data row with the corresponding prediction resultrequests_exploded = (requests_with_features.withColumn("feature_prediction_pairs",F.arrays_zip(F.col("feature_data"), F.col("predictions"))).withColumn("feature_prediction_pairs",F.explode(F.col("feature_prediction_pairs"))).withColumn("feature_row", F.col("feature_prediction_pairs.feature_data")).withColumn(PREDICTION_COL, F.col("feature_prediction_pairs.predictions")))# 3. dynamically create feature columnrequests_with_feature_cols = requests_explodedfor i, col_name in enumerate(FEATURE_COLUMNS):requests_with_feature_cols = (requests_with_feature_cols.withColumn(col_name, F.col("feature_row")[i]))# 4. Add record unique ID, making it easy to perform upsert operationrequests_with_example_id = requests_with_feature_cols \\.withColumn(EXAMPLE_ID_COL, F.expr("uuid()"))# 5. Clean up temporary columnsrequests_processed = (requests_with_example_id.drop("feature_columns", "feature_data", "feature_prediction_pairs", "feature_row", "predictions", "request", "response"))return requests_processeddef create_table(spark: SparkSession,table_name: str,df: Optional[DataFrame] = None,partition_expr: str = None,description: Optional[str] = None,):"""Create a Hive table.Args:spark: SparkSession instancetable_name: full hive table name (format: database.data table)df: initial data (optional, used for infer schema)partition_expr: partition column (optimized storage query)description: table descriptionRaises:ValueError: throw create exception"""# Infer table schematable_schema = df.schemapart_col_nam = partition_expr.split(' ')[0]# Build column definitioncolumns_ddl = []for field in table_schema.fields:# Filter out partition columns when creating a tableif field.name == part_col_nam:continuedata_type = field.dataType.simpleString().upper()col_def = f"`{field.name}` {data_type}"if not field.nullable:col_def += " NOT NULL"columns_ddl.append(col_def)# Build table creation statementddl = f"""CREATE TABLE IF NOT EXISTS {table_name} ({', '.join(columns_ddl)})PARTITIONED BY ({partition_expr})TBLPROPERTIES ('comment'= '{description or ''}')"""# Print sqlprint(f"create table ddl: {ddl}\\n")# execute DDLtry:spark.sql(ddl)except Exception as e:raise ValueError(f"Failed to create table: {str(e)}") from eprint(f"create table {table_name} done")def write_batch_to_hive(batch_df: DataFrame, batch_id: int, table_name: str):# Control small files: repartition by partition columnout_df = batch_df.repartition(8, "dt")out_df.write.mode("append").format("hive").saveAsTable(table_name,partitionBy="dt")spark = SparkSession.builder.appName("Operate DB Example").enableHiveSupport().getOrCreate()# Build a streaming DataFramedf = (spark.readStream.format("json").schema(schema).load(SOURCE_PATH))# Process streaming dataunpacked_df = process_requests(df)# Insert partition columnrequests_with_dt = (unpacked_df.withColumn("dt", F.substring(F.col("request_date"), 1, 10)) # Extract date in YYYY-MM-DD format)# Create a result tablecreate_table(spark, UNPACKED_TABLE_NAME, unpacked_df, 'dt STRING')# Set Hive dynamic partitionspark.sql("SET hive.exec.dynamic.partition=true")spark.sql("SET hive.exec.dynamic.partition.mode=nonstrict")# Batch write to result tablequery = (requests_with_dt.writeStream.outputMode("append").option("checkpointLocation", CHECKPOINT_PATH).trigger(once=True).foreachBatch(lambda df, id: write_batch_to_hive(df, id, UNPACKED_TABLE_NAME)).start())query.awaitTermination()# Query result dataspark.sql(f"select * from {UNPACKED_TABLE_NAME}").show(10)
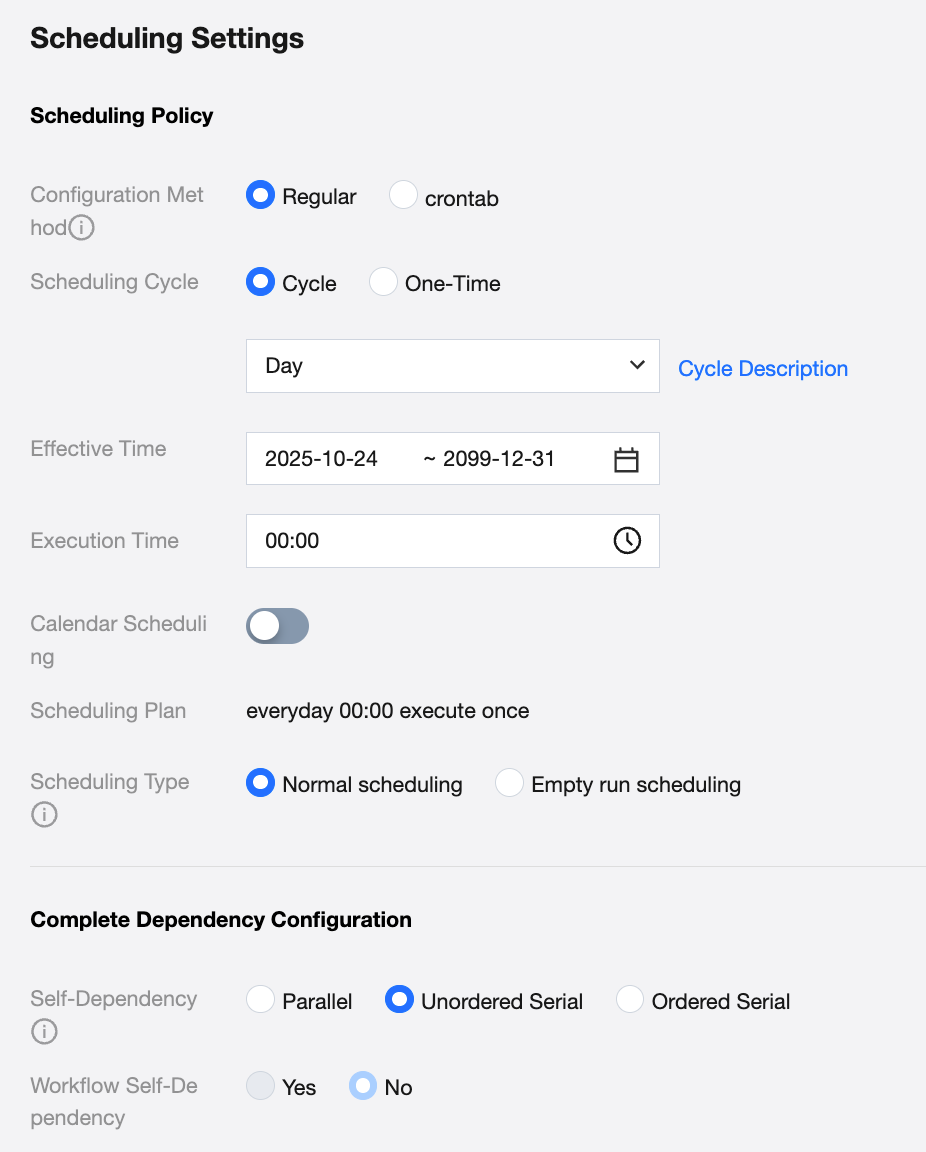
Configure Task Cycle Operation
Debug the task, ensure manual execution successful and write the processed data, then you can configure the task as scheduled execution in scheduling settings. Recommended configuration is at least once a week to continuously monitor model quality.
Configure Association Tag Data Task
To build a complete model monitoring and evaluation system, it is necessary to associate these inference data with actual tag data, forming a comprehensive inference table dataset that includes input features, model prediction results, and true labels. This tag association process is a key step in model effectiveness evaluation. By comparing predicted values with true labels, core performance metrics such as accuracy and recall rate can be calculated, enabling quantitative monitoring and trend analysis of model quality. Obtaining tag data typically requires integrating feedback mechanisms from business systems or manual annotation processes. Depending on the data processing engine and specific business scenarios, you can refer to the corresponding data processing code implementation to complete tag association operations, establishing a complete data foundation for comprehensive model monitoring.
DLC Spark Engine
from pyspark.sql import DataFrame, SparkSessionfrom typing import Union, List, Dict, Optional, Sequence, Any# Table name for the reasoning table to be associated with a tag: It needs to be a three-segment table name in DLC: <data catalog>.<database>.<data table>UNPACKED_TABLE_NAME = "DataLakeCatalog.test.unpacked_test_inference_table"# Tag data table configuration: can be multiple, in each table must contain structure (<table name>, <list of field names to be retained in join>, <list of field names used for equivalent condition connection in join>)JOIN_TABLES = [('DataLakeCatalog.test.test_label_table', ['label', 'example_id'], ['example_id'])]# Table name for reasoning after associating tag dataFULLY_QUALIFIED_TABLE_NAME = "DataLakeCatalog.test.test_fully_qualified_table_name"# Window size: Limits how long the data processed by this Notebook script each run. Data older than this window will be ignored if not yet processed. To ensure all data is processed, the scheduling interval should be smaller than this window size.PROCESSING_WINDOW_DAYS = 10# primary key field for result table insertionMERGE_COLS = ["request_time", "example_id"]def create_table(spark: SparkSession,table_name: str,df: Optional[DataFrame] = None,partition_expr: str = None,description: Optional[str] = None,):"""Create a dlc internal tableArgs:spark: SparkSession instancetable_name: full table name (format: data catalog.database.data table)df: initial data (used for infer schema)partition_expr: partition column (optimized storage query)description: table descriptionRaises:ValueError: throw create exception"""# Infer table schematable_schema = df.schema# Build column definitioncolumns_ddl = []for field in table_schema.fields:data_type = field.dataType.simpleString().upper()col_def = f"`{field.name}` {data_type}"if not field.nullable:col_def += " NOT NULL"columns_ddl.append(col_def)# Build table creation statementddl = f"""CREATE TABLE IF NOT EXISTS {table_name} ({', '.join(columns_ddl)})USING icebergPARTITIONED BY ({partition_expr})TBLPROPERTIES ('comment'= '{description or ''}','format-version'= '2','write.metadata.previous-versions-max'= '100','write.metadata.delete-after-commit.enabled'= 'true','smart-optimizer.inherit' = 'none','smart-optimizer.written.enable' = 'enable')"""# Print sqlprint(f"create table ddl: {ddl}\\n")# execute DDLtry:spark.sql(ddl)except Exception as e:raise ValueError(f"Failed to create table: {str(e)}") from eprint(f"create table {table_name} done")def upsert_table(spark: SparkSession,target_table_name: str,merge_cols: list,df: DataFrame,):"""Write data to the target table using the upsert methodArgs:spark: SparkSession instancetarget_table_name: full result table name (format: data catalog.database.data table)merge_cols: primary key when updatingdf: data to be updatedRaises:ValueError: throw write exception"""merge_condition = " AND ".join([f"target.{col} = source.{col}" for col in merge_cols])# Create a temporary viewdf.createOrReplaceTempView("source_data")# Perform MERGE operationmerge_sql = f"""MERGE INTO {target_table_name} AS targetUSING source_data AS sourceON {merge_condition}WHEN MATCHED THEN UPDATE SET *WHEN NOT MATCHED THEN INSERT *"""# execute DMLtry:spark.sql(merge_sql)except Exception as e:raise ValueError(f"Failed to create table: {str(e)}") from eprint(f"create table {table_name} done")spark = SparkSession.builder.appName("Operate DB Example").getOrCreate()# read data of PROCESSING_WINDOW_DAYS each timerequests_processed = spark.table(UNPACKED_TABLE_NAME) \\.filter(f"CAST(request_time AS DATE) >= current_date() - (INTERVAL {PROCESSING_WINDOW_DAYS} DAYS)")if requests_processed.count() > 0:# Associated tag tablefor table_name, preserve_cols, join_cols in JOIN_TABLES:join_data = spark.table(table_name)requests_processed = requests_processed.join(join_data.select(preserve_cols), on=join_cols, how="left")# Create a result tablecreate_table(spark, FULLY_QUALIFIED_TABLE_NAME, requests_processed, 'days(request_time)')# Write to result tableupsert_table(spark, FULLY_QUALIFIED_TABLE_NAME, MERGE_COLS, requests_processed)spark.sql(f"select * from {FULLY_QUALIFIED_TABLE_NAME}").show(10)
EMR Spark Engine
from pyspark.sql import DataFrame, SparkSessionfrom typing import Union, List, Dict, Optional, Sequence, Any# Name of the table to be associated with tags: needs to be a Hive table name: <database>.<data table>UNPACKED_TABLE_NAME = "testdb.unpacked_test_inference_table"# Tag data table configuration: can be multiple, in each table must contain structure (<table name>, <list of field names to be retained in join>, <list of field names used for equivalent condition connection in join>)JOIN_TABLES = [('testdb.test_label_table', ['label', 'example_id'], ['example_id'])]# Table name for reasoning after associating tag dataFULLY_QUALIFIED_TABLE_NAME = "testdb.test_fully_qualified_table_name"# Window size: Limits how long the data processed by this Notebook script each run. Data older than this window will be ignored if not yet processed. To ensure all data is processed, the scheduling interval should be smaller than this window size.PROCESSING_WINDOW_DAYS = 10def create_table(spark: SparkSession,table_name: str,df: Optional[DataFrame] = None,partition_expr: str = None,description: Optional[str] = None,):"""Create a Hive table.Args:spark: SparkSession instancetable_name: full hive table name (format: database.data table)df: initial data (optional, used for infer schema)partition_expr: partition column (optimized storage query)description: table descriptionRaises:ValueError: throw create exception"""# Infer table schematable_schema = df.schemapart_col_nam = partition_expr.split(' ')[0]# Build column definitioncolumns_ddl = []for field in table_schema.fields:# Filter out partition columns when creating a tableif field.name == part_col_nam:continuedata_type = field.dataType.simpleString().upper()col_def = f"`{field.name}` {data_type}"if not field.nullable:col_def += " NOT NULL"columns_ddl.append(col_def)# Build table creation statementddl = f"""CREATE TABLE IF NOT EXISTS {table_name} ({', '.join(columns_ddl)})PARTITIONED BY ({partition_expr})TBLPROPERTIES ('comment'= '{description or ''}')"""# Print sqlprint(f"create table ddl: {ddl}\\n")# execute DDLtry:spark.sql(ddl)except Exception as e:raise ValueError(f"Failed to create table: {str(e)}") from eprint(f"create table {table_name} done")def partition_overwrite(spark, source_df, target_table):""""""try:Check if data is emptyif source_df.count() == 0:print(f"Warning: no data writing required to {target_table}")return# Display partitions to overwritepartitions = source_df.select("dt").distinct().collect()partition_list = [row.dt for row in partitions]print(f"Partitions to overwrite: {partition_list}")# Set parameters and writespark.sql("SET hive.exec.dynamic.partition=true")spark.sql("SET hive.exec.dynamic.partition.mode=nonstrict")spark.conf.set("spark.sql.sources.partitionOverwriteMode", "dynamic")source_df.repartition(8, "dt") \\.write \\.mode("overwrite") \\.format("hive") \\.saveAsTable(target_table)print(f"Successfully overwrote {len(partition_list)} partitions to table {target_table}")except Exception as e:print(f"Error occurred while writing to table {target_table}: {str(e)}")raisespark = SparkSession.builder.appName("Operate DB Example").getOrCreate()# read data of PROCESSING_WINDOW_DAYS partition each time, then overwrite after processingrequests_processed = spark.table(UNPACKED_TABLE_NAME) \\.filter(f"dt >= date_format(current_date() - INTERVAL {PROCESSING_WINDOW_DAYS} DAYS, 'yyyy-MM-dd')")for table_name, preserve_cols, join_cols in JOIN_TABLES:join_data = spark.table(table_name)requests_processed = requests_processed.join(join_data.select(preserve_cols), on=join_cols, how="left")create_table(spark, FULLY_QUALIFIED_TABLE_NAME, requests_processed, 'dt STRING')partition_overwrite(spark, requests_processed, FULLY_QUALIFIED_TABLE_NAM)spark.sql(f"select * from {FULLY_QUALIFIED_TABLE_NAME}").show(10)
Configure Task Cycle Operation
After ensuring the tag task is successfully executed manually and data is written, you can configure the task for periodic execution. For the setting method, refer to procedure 2. The recommended performance period should be equal to or greater than the configured COS data processing task period in procedure 2.
Creating a Model Service Resource Group (CVM Resource Group)
When deploying a model service, you need to configure the resource source. Select CVM from the dropdown list to display the available resource group for the current project. You can freely configure the amount of CPU and memory, and toggle GPU usage.
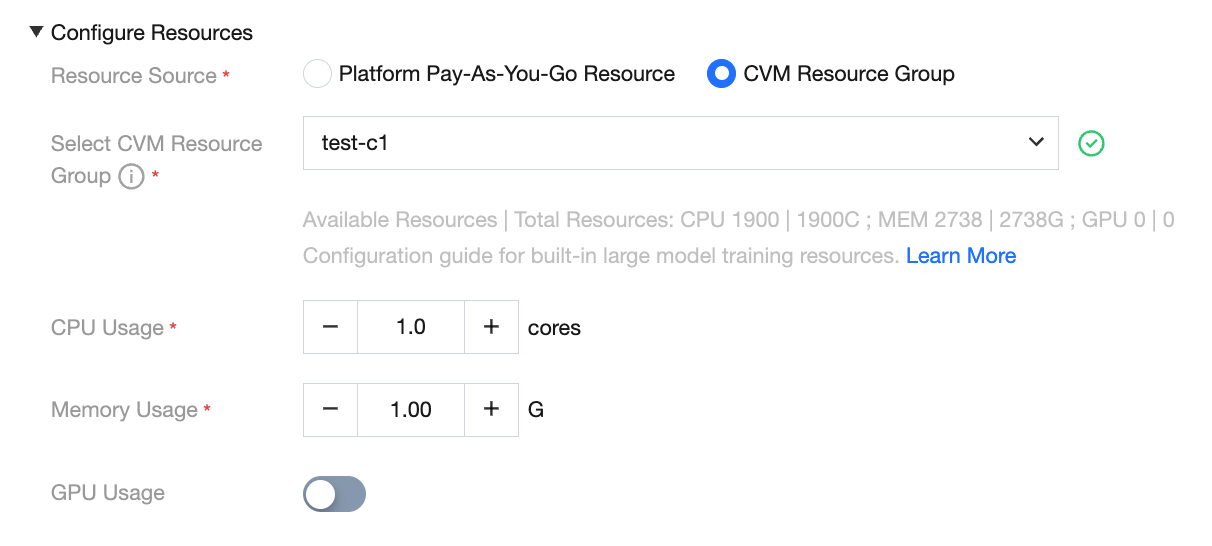
Click Manage CVM resource group to navigate to the resource group list page.
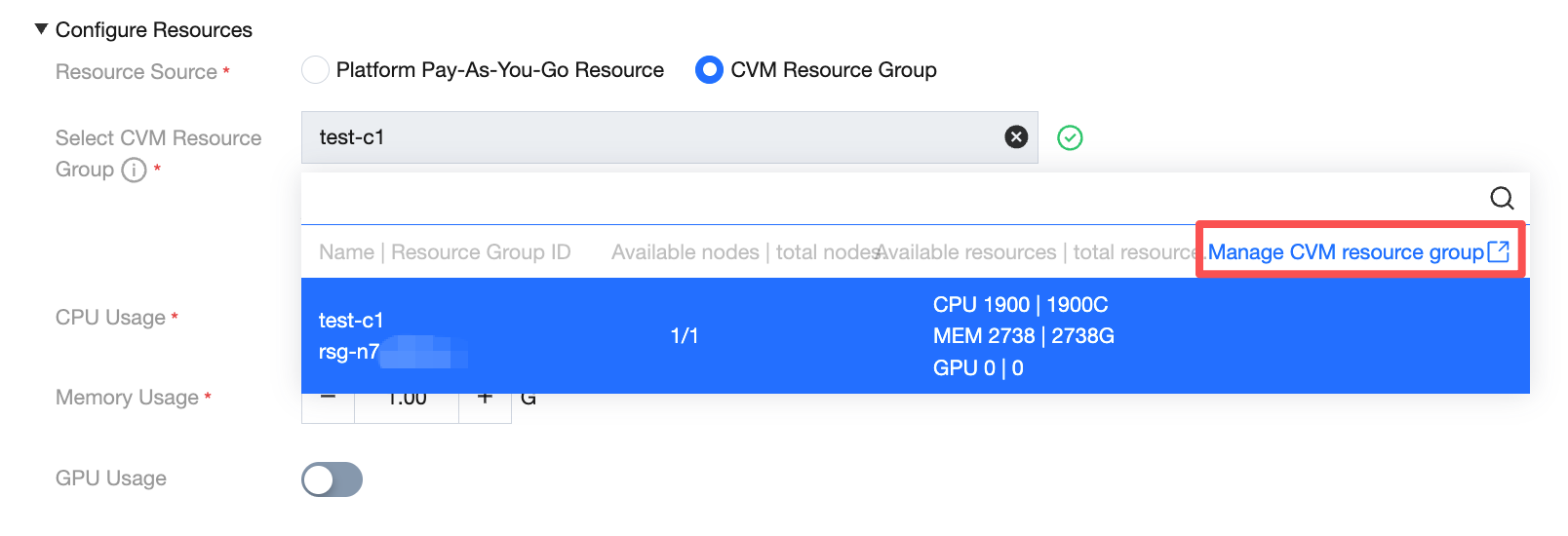
View model service - resource group list, click the action button to associate with a project.
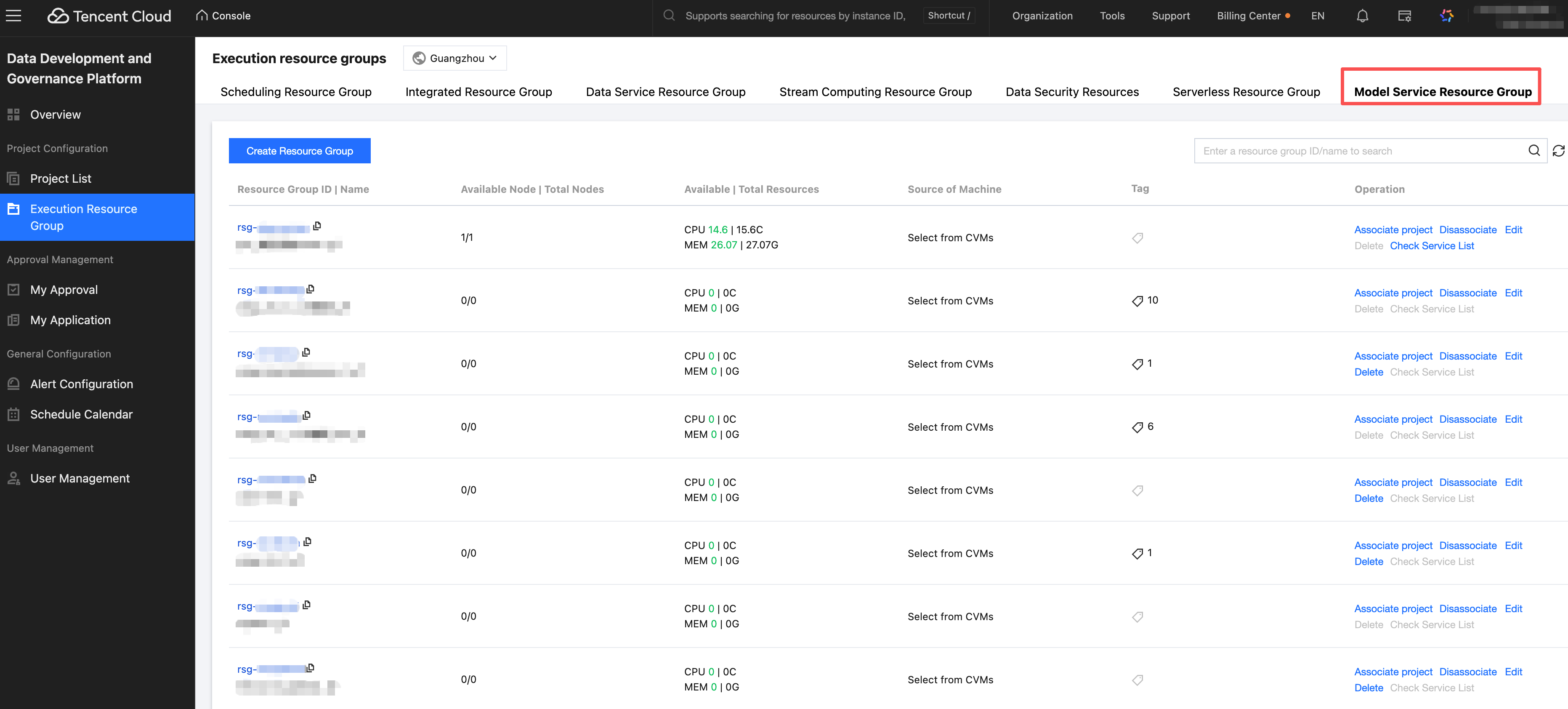
Select operation - Associate project button, you can filter the project name from the drop-down and complete the association.
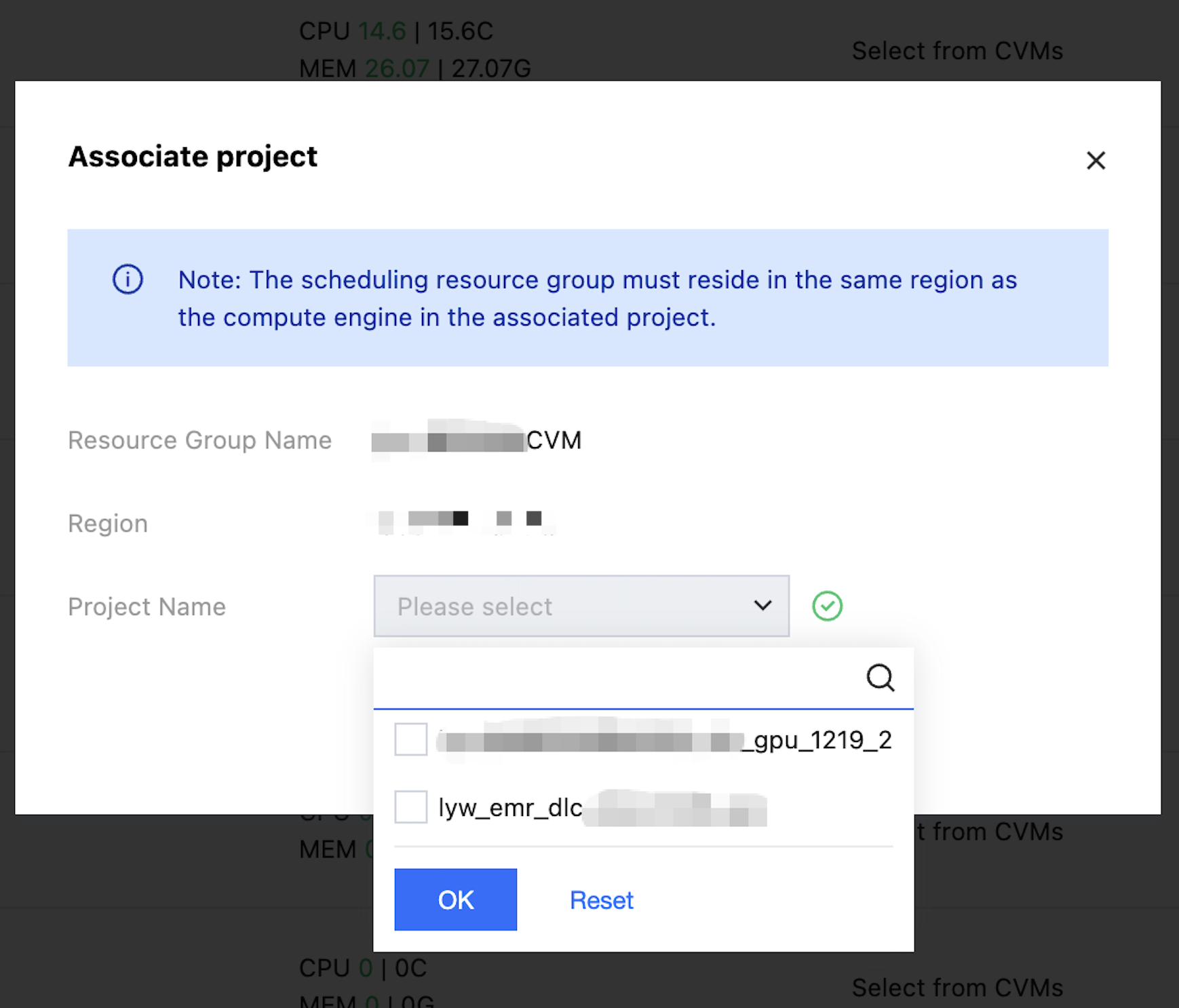
Click Unbind to disassociate the project from the resource group.
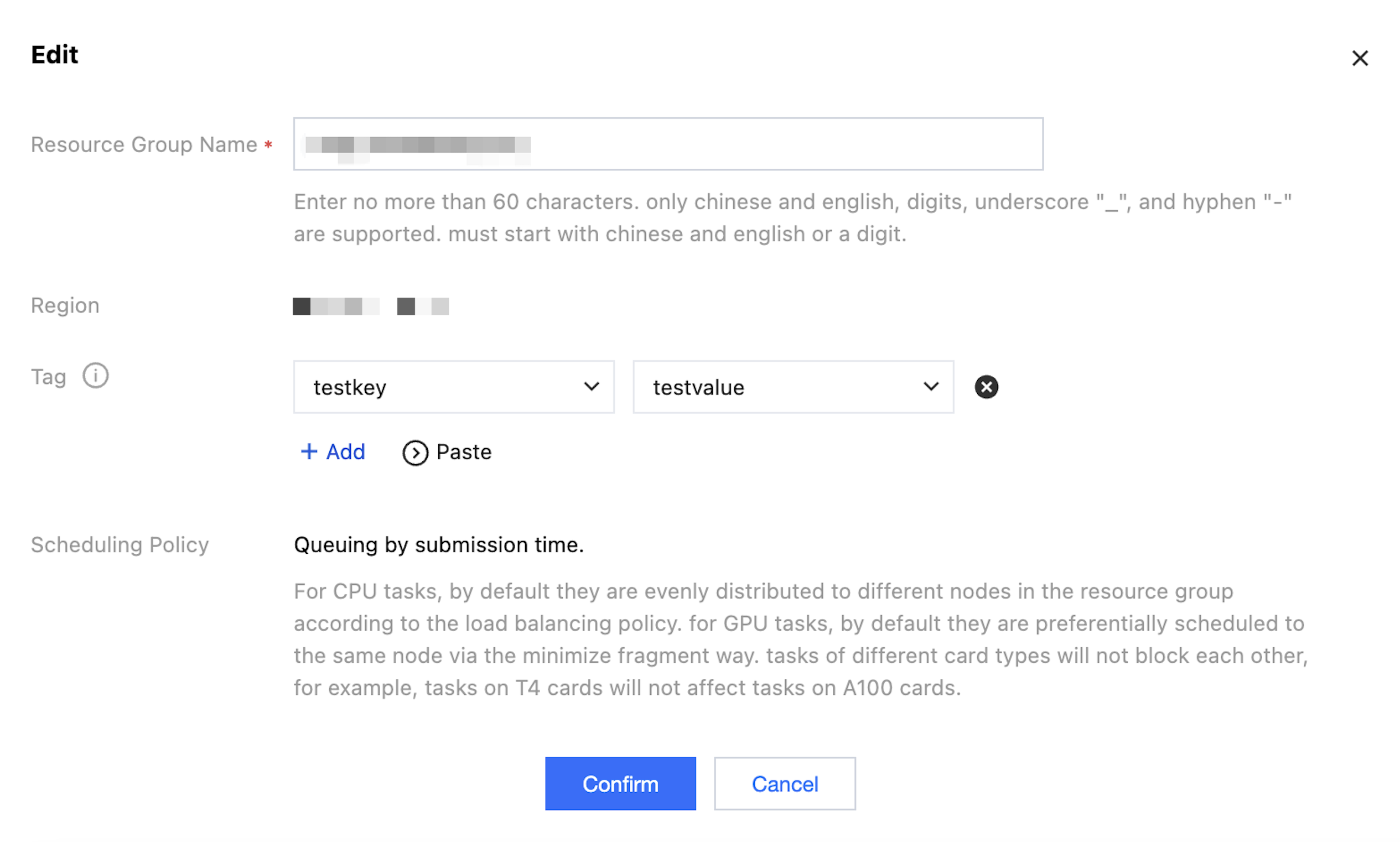
Click Edit to modify the tag, name, and other fields of the resource group.
Click the resource group name to enter the resource group details page.
You can view node ID, available resources, running state, billing mode, time, etc. It supports checking the service list, moving, renewing, isolating, and releasing.
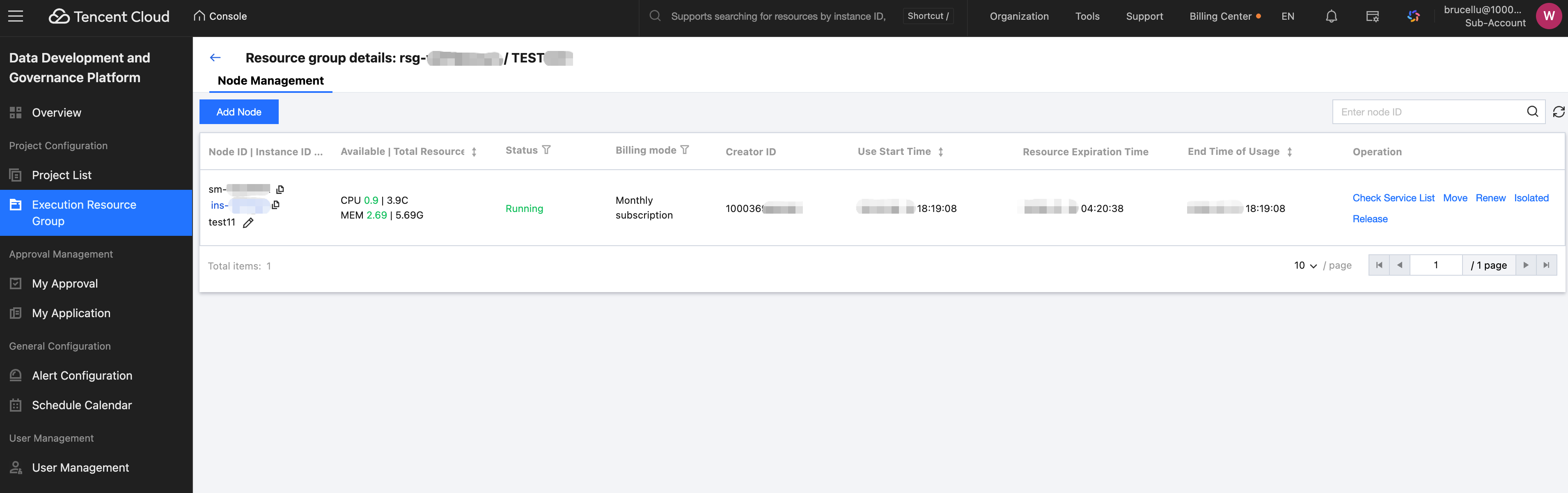
Click Add Node to navigate to the payment page.
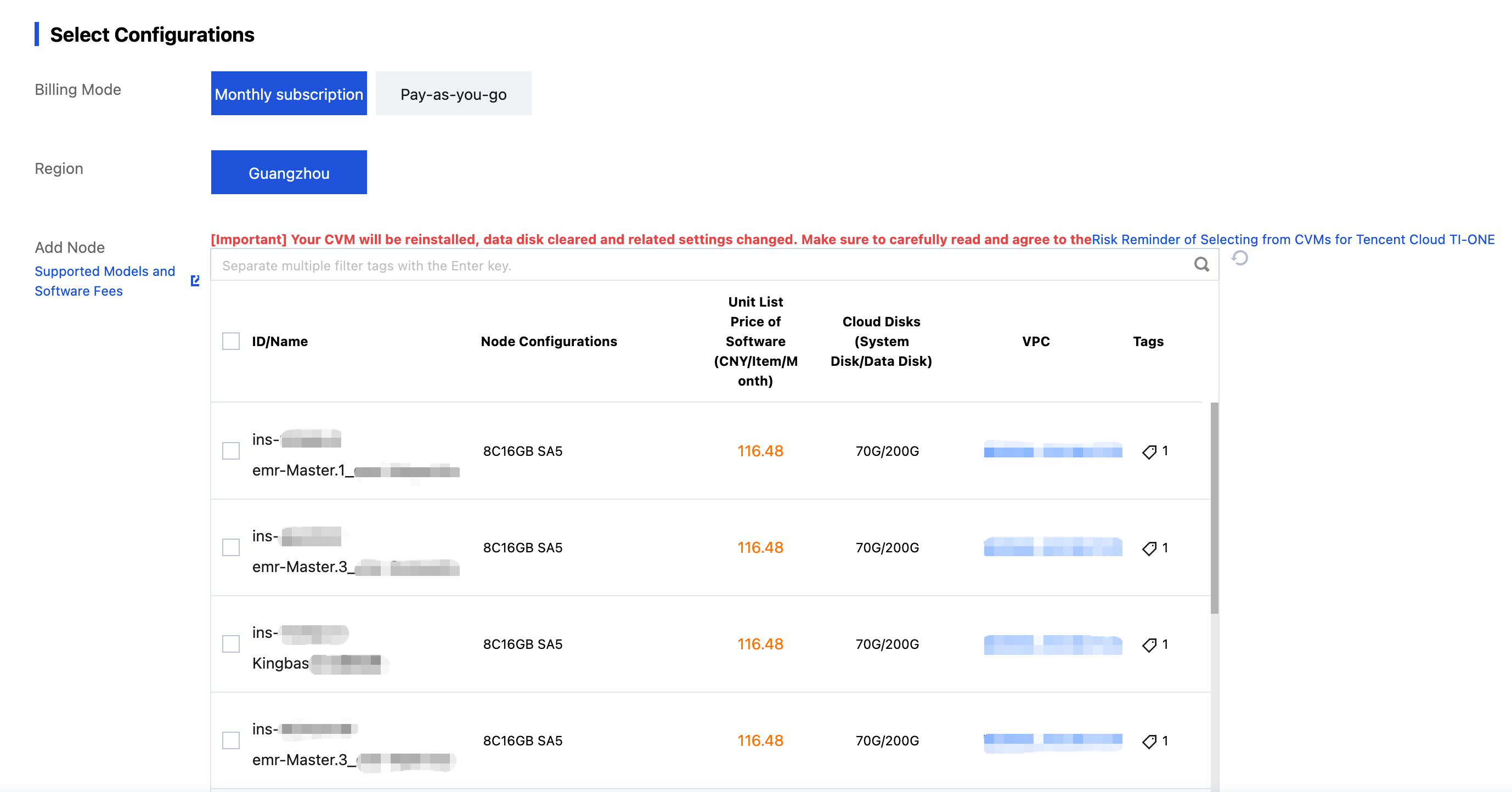
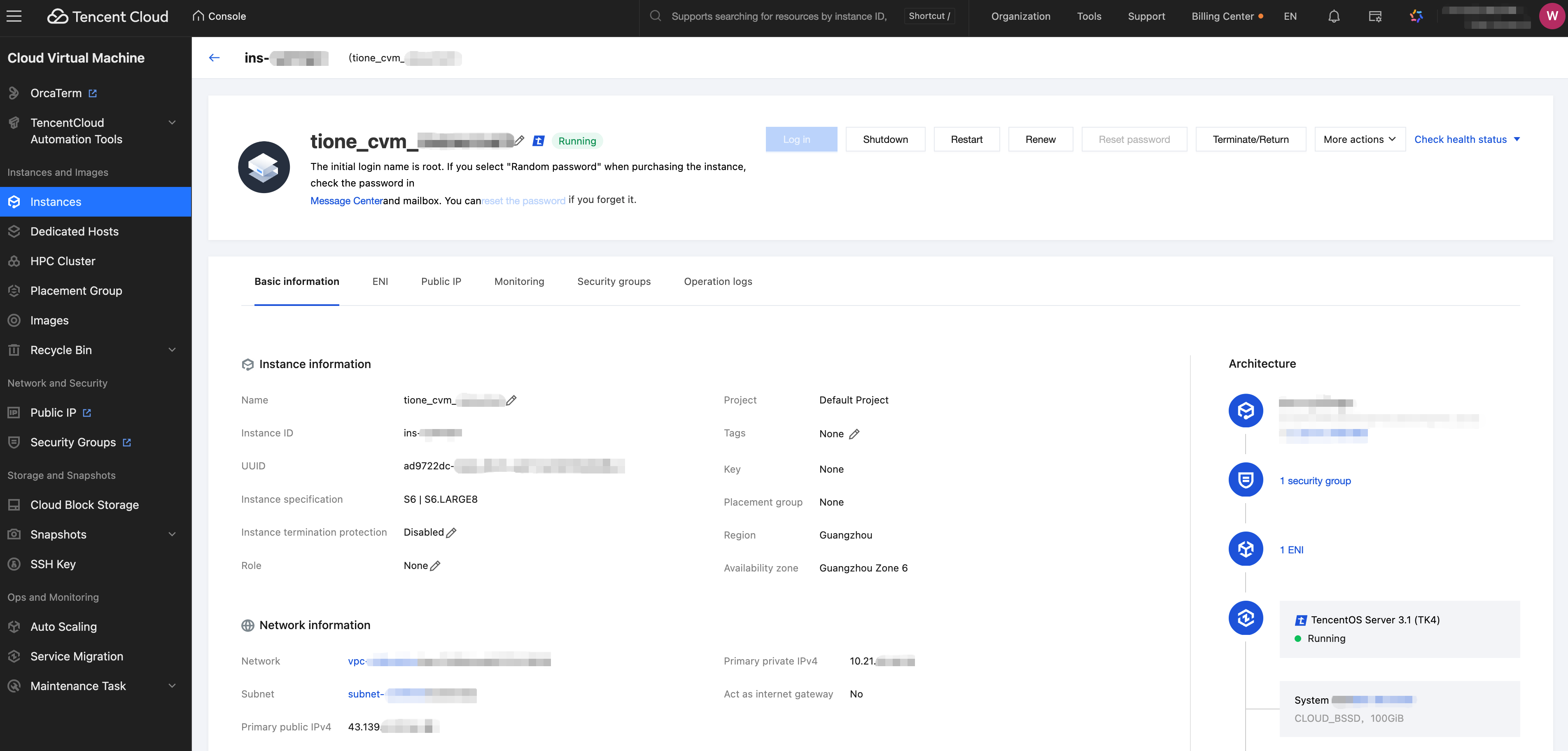
Click the instance ID to navigate to the instance details page. You can log in, power off, restart, renew, reset the password, terminate/return the instance, and show instance-related information.
Help and Support
Was this page helpful?
You can also Contact sales or Submit a Ticket for help.
Help us improve! Rate your documentation experience in 5 mins.
Feedback